-
Evans Syndrome
Wikipedia
"Primary thrombocytopenic purpura and acquired hemolytic anemia; evidence for a common etiology". Archives of Internal Medicine . 87 (1): 48–65. doi : 10.1001/archinte.1951.03810010058005 . ... "Evans syndrome in adults - incidence, prevalence, and survival in a nationwide cohort". Am J Hematol . 94 (10): 1081–1090. doi : 10.1002/ajh.25574 . ... Retrieved 2019-07-28 . ^ Pegels JG, Helmerhorst FM, van Leeuwen EF, van de Plas-van Dalen C, Engelfriet CP, von dem Borne AE (1982). ... "Evans syndrome in childhood". J. Pediatr . 97 (5): 754–8. doi : 10.1016/S0022-3476(80)80258-7 . ... PMID 9313889 . ^ Oyama Y, Papadopoulos EB, Miranda M, Traynor AE, Burt RK (2001). "Allogeneic stem cell transplantation for Evans syndrome" .
-
Partial Androgen Insensitivity Syndrome
Wikipedia
Some individuals with PAIS have a sufficiently high sperm count to father children; at least one case report has been published that describes fertile men who fit the criteria for grade 2 PAIS ( micropenis , penile hypospadias , and gynecomastia ). [36] Several publications have indicated that testosterone treatment can correct low sperm counts in men with MAIS. [1] [78] At least one case report has been published that documents the efficacy of treating a low sperm-count with tamoxifen in an individual with PAIS. [81] Counseling [ edit ] Depending on phenotypic features, impotence and other sexual problems such as anejaculation or sexual aversion may be fairly common among individuals with PAIS, [20] [28] [29] [30] [31] but do not necessarily indicate low libido . [26] [28] Support groups for individuals with PAIS may help affected individuals discuss their concerns more comfortably. [26] Some individuals with PAIS may try to avoid intimate relationships out of fear of rejection; individual therapy may help some to overcome social anxiety , and restore focus to interpersonal relationships instead of solely on sexual function and activity. [26] Society and culture [ edit ] Adults with partial androgen insensitivity syndrome include Australian-Maltese advocate Tony Briffa , considered to be the world's first openly intersex mayor and public office-bearer. [82] Briffa served as Deputy Mayor of the City of Hobsons Bay, Victoria, between 2009 and 2011, and Mayor between 2011–2012. [83] [82] [84] [85] [86] [87] In history , the Roman sophist and philosopher Favorinus of Arelate has been described as having partial androgen insensitivity syndrome. [88] [89] Notable people with PAIS [ edit ] Tony Briffa [90] [91] Small Luk [92] Eliana Rubashkyn [93] [94] [95] Sean Saifa Wall [96] Sentencia SU 337/99, Colombia [ edit ] Further information: Intersex rights in Colombia In Sentencia SU-337/99, of May 12, 1999, the Constitutional Court of Colombia determined that "qualified and persistent" informed consent is required for genital surgeries in children. ... Clin. Endocrinol. Metab . 20 (4): 577–98. doi : 10.1016/j.beem.2006.11.003 . ... "A novel mutation c.118delA in exon 1 of the androgen receptor gene resulting in complete androgen insensitivity syndrome within a large family". Fertil. Steril . 89 (5): 1260.e3–7. doi : 10.1016/j.fertnstert.2007.04.057 . ... PMID 16757528 . ^ a b c d e Melo KF, Mendonca BB, Billerbeck AE, Costa EM, Inácio M, Silva FA, Leal AM, Latronico AC, Arnhold IJ (July 2003). ... Lancet . 344 (8925): 826–7. doi : 10.1016/S0140-6736(94)92385-X . PMID 7993455 . ^ a b Nieschlag E (September 2006).
-
Border Disease
Wikipedia
Viral disease of sheep and goats caused by Pestivirus D Pestivirus D Virus classification (unranked): Virus Realm : Riboviria Kingdom: Orthornavirae Phylum: Kitrinoviricota Class: Flasuviricetes Order: Amarillovirales Family: Flaviviridae Genus: Pestivirus Species: Pestivirus D Synonyms . [1] Border disease virus Border disease ( BD ) is a viral disease of sheep and goats , primarily causing congenital diseases, but can also cause acute and persistent infections. It first appeared in the border regions of England and Wales in 1959, and has since spread world-wide. Lambs that are born with BD are commonly known as 'hairy shakers' due to the primary presentation of the disease.
-
Congenital Hypothyroidism
Wikipedia
Persistence of severe, untreated hypothyroidism resulted in severe mental impairment, with an IQ below 80 in the majority. Most of these children eventually ended up in institutional care. [ citation needed ] 3 month old infant with untreated CH; picture demonstrates hypotonic posture, myxedematous facies, macroglossia, and umbilical hernia Close up of face, showing myxedematous facies, macroglossia, and skin mottling Close up showing abdominal distension and umbilical hernia. ... Adaptive behavior Fine motor Gross motor Language Personal-social behavior Severe CH 92 89 90 89 90 Moderate CH 97 97 98 96 96 Mild CH 100 99 100 99 100 [8] Epidemiology [ edit ] Congenital hypothyroidism (CH) occurs in 1:1300 to 1:4000 births worldwide. [9] [10] [11] [12] [13] The differences in CH-incidence are more likely due to iodine deficiency thyroid disorders or to the type of screening method than to ethnic affiliation. [10] CH is caused by an absent or defective thyroid gland classified into agenesis (22-42%), ectopy (35-42%) and gland in place defects (24-36%). [10] [14] It is also found to be of increased association with female sex and gestational age >40 weeks. [14] References [ edit ] ^ Worth, Chris; Hird, Beverly; Tetlow, Lesley; Wright, Neville; Patel, Leena; Banerjee, Indraneel (14 November 2019). ... Molecular Genetics and Metabolism . 91 (3): 268–77. doi : 10.1016/j.ymgme.2007.03.012 . ... External links [ edit ] Classification D ICD - 10 : E00 , E03.0 , E03.1 ICD - 9-CM : 243 MeSH : D003409 DiseasesDB : 6612 External resources MedlinePlus : 001193 v t e Thyroid disease Hypothyroidism Iodine deficiency Cretinism Congenital hypothyroidism Myxedema Myxedema coma Euthyroid sick syndrome Signs and symptoms Queen Anne's sign Woltman sign Thyroid dyshormonogenesis Pickardt syndrome Hyperthyroidism Hyperthyroxinemia Thyroid hormone resistance Familial dysalbuminemic hyperthyroxinemia Hashitoxicosis Thyrotoxicosis factitia Thyroid storm Graves' disease Signs and symptoms Abadie's sign of exophthalmic goiter Boston's sign Dalrymple's sign Stellwag's sign lid lag Griffith's sign Möbius sign Pretibial myxedema Graves' ophthalmopathy Thyroiditis Acute infectious Subacute De Quervain's Subacute lymphocytic Palpation Autoimmune /chronic Hashimoto's Postpartum Riedel's Enlargement Goitre Endemic goitre Toxic nodular goitre Toxic multinodular goiter Thyroid nodule Colloid nodule v t e Congenital endocrine disorders Pituitary Congenital hypopituitarism Thyroid Thyroid disease Persistent thyroglossal duct Thyroglossal cyst Congenital hypothyroidism Thyroid dysgenesis Thyroid dyshormonogenesis Pendred syndrome Parathyroid Congenital absence of parathyroid Adrenal Absent adrenal gland v t e Cell surface receptor deficiencies G protein-coupled receptor (including hormone ) Class A TSHR ( Congenital hypothyroidism 1 ) LHCGR ( Luteinizing hormone insensitivity , Leydig cell hypoplasia , Male-limited precocious puberty ) FSHR ( Follicle-stimulating hormone insensitivity , XX gonadal dysgenesis ) GnRHR ( Gonadotropin-releasing hormone insensitivity ) EDNRB ( ABCD syndrome , Waardenburg syndrome 4a , Hirschsprung's disease 2 ) AVPR2 ( Nephrogenic diabetes insipidus 1 ) PTGER2 ( Aspirin-induced asthma ) Class B PTH1R ( Jansen's metaphyseal chondrodysplasia ) Class C CASR ( Familial hypocalciuric hypercalcemia ) Class F FZD4 ( Familial exudative vitreoretinopathy 1 ) Enzyme-linked receptor (including growth factor ) RTK ROR2 ( Robinow syndrome ) FGFR1 ( Pfeiffer syndrome , KAL2 Kallmann syndrome ) FGFR2 ( Apert syndrome , Antley–Bixler syndrome , Pfeiffer syndrome , Crouzon syndrome , Jackson–Weiss syndrome ) FGFR3 ( Achondroplasia , Hypochondroplasia , Thanatophoric dysplasia , Muenke syndrome ) INSR ( Donohue syndrome Rabson–Mendenhall syndrome ) NTRK1 ( Congenital insensitivity to pain with anhidrosis ) KIT ( KIT Piebaldism , Gastrointestinal stromal tumor ) STPK AMHR2 ( Persistent Müllerian duct syndrome II ) TGF beta receptors : Endoglin / Alk-1 / SMAD4 ( Hereditary hemorrhagic telangiectasia ) TGFBR1 / TGFBR2 ( Loeys–Dietz syndrome ) GC GUCY2D ( Leber's congenital amaurosis 1 ) JAK-STAT Type I cytokine receptor : GH ( Laron syndrome ) CSF2RA ( Surfactant metabolism dysfunction 4 ) MPL ( Congenital amegakaryocytic thrombocytopenia ) TNF receptor TNFRSF1A ( TNF receptor associated periodic syndrome ) TNFRSF13B ( Selective immunoglobulin A deficiency 2 ) TNFRSF5 ( Hyper-IgM syndrome type 3 ) TNFRSF13C ( CVID4 ) TNFRSF13B ( CVID2 ) TNFRSF6 ( Autoimmune lymphoproliferative syndrome 1A ) Lipid receptor LRP : LRP2 ( Donnai–Barrow syndrome ) LRP4 ( Cenani–Lenz syndactylism ) LRP5 ( Worth syndrome , Familial exudative vitreoretinopathy 4 , Osteopetrosis 1 ) LDLR ( LDLR Familial hypercholesterolemia ) Other/ungrouped Immunoglobulin superfamily : AGM3, 6 Integrin : LAD1 Glanzmann's thrombasthenia Junctional epidermolysis bullosa with pyloric atresia EDAR ( EDAR hypohidrotic ectodermal dysplasia ) PTCH1 ( Nevoid basal-cell carcinoma syndrome ) BMPR1A ( BMPR1A juvenile polyposis syndrome ) IL2RG ( X-linked severe combined immunodeficiency ) See also cell surface receptors v t e Genetic disorders relating to deficiencies of transcription factor or coregulators (1) Basic domains 1.2 Feingold syndrome Saethre–Chotzen syndrome 1.3 Tietz syndrome (2) Zinc finger DNA-binding domains 2.1 ( Intracellular receptor ): Thyroid hormone resistance Androgen insensitivity syndrome PAIS MAIS CAIS Kennedy's disease PHA1AD pseudohypoaldosteronism Estrogen insensitivity syndrome X-linked adrenal hypoplasia congenita MODY 1 Familial partial lipodystrophy 3 SF1 XY gonadal dysgenesis 2.2 Barakat syndrome Tricho–rhino–phalangeal syndrome 2.3 Greig cephalopolysyndactyly syndrome / Pallister–Hall syndrome Denys–Drash syndrome Duane-radial ray syndrome MODY 7 MRX 89 Townes–Brocks syndrome Acrocallosal syndrome Myotonic dystrophy 2 2.5 Autoimmune polyendocrine syndrome type 1 (3) Helix-turn-helix domains 3.1 ARX Ohtahara syndrome Lissencephaly X2 MNX1 Currarino syndrome HOXD13 SPD1 synpolydactyly PDX1 MODY 4 LMX1B Nail–patella syndrome MSX1 Tooth and nail syndrome OFC5 PITX2 Axenfeld syndrome 1 POU4F3 DFNA15 POU3F4 DFNX2 ZEB1 Posterior polymorphous corneal dystrophy Fuchs' dystrophy 3 ZEB2 Mowat–Wilson syndrome 3.2 PAX2 Papillorenal syndrome PAX3 Waardenburg syndrome 1&3 PAX4 MODY 9 PAX6 Gillespie syndrome Coloboma of optic nerve PAX8 Congenital hypothyroidism 2 PAX9 STHAG3 3.3 FOXC1 Axenfeld syndrome 3 Iridogoniodysgenesis, dominant type FOXC2 Lymphedema–distichiasis syndrome FOXE1 Bamforth–Lazarus syndrome FOXE3 Anterior segment mesenchymal dysgenesis FOXF1 ACD/MPV FOXI1 Enlarged vestibular aqueduct FOXL2 Premature ovarian failure 3 FOXP3 IPEX 3.5 IRF6 Van der Woude syndrome Popliteal pterygium syndrome (4) β-Scaffold factors with minor groove contacts 4.2 Hyperimmunoglobulin E syndrome 4.3 Holt–Oram syndrome Li–Fraumeni syndrome Ulnar–mammary syndrome 4.7 Campomelic dysplasia MODY 3 MODY 5 SF1 SRY XY gonadal dysgenesis Premature ovarian failure 7 SOX10 Waardenburg syndrome 4c Yemenite deaf-blind hypopigmentation syndrome 4.11 Cleidocranial dysostosis (0) Other transcription factors 0.6 Kabuki syndrome Ungrouped TCF4 Pitt–Hopkins syndrome ZFP57 TNDM1 TP63 Rapp–Hodgkin syndrome / Hay–Wells syndrome / Ectrodactyly–ectodermal dysplasia–cleft syndrome 3 / Limb–mammary syndrome / OFC8 Transcription coregulators Coactivator: CREBBP Rubinstein–Taybi syndrome Corepressor: HR ( Atrichia with papular lesions ) v t e Extracellular ligand disorders Cytokine EDA Hypohidrotic ectodermal dysplasia Camurati–Engelmann disease Ephrin Craniofrontonasal dysplasia WNT Tetra-amelia syndrome TGF OFC 11 Fas ligand Autoimmune lymphoproliferative syndrome 1B Endothelin EDN3 Waardenburg syndrome IVb Hirschsprung's disease 4 Other DHH ( DHH XY gonadal dysgenesis ) BMP15 ( Premature ovarian failure 4 ) TSHB ( Congenital hypothyroidism 4 ) See also intercellular signaling peptides and proteinsTSHR, DUOX2, TPO, PAX8, FOXE1, TG, GLIS3, NKX2-1, IGSF1, TSHB, IYD, POU1F1, CDCA8, TRHR, INHBB, ATP5PD, ARPC5, IGF1, NGFR, NEFL, PPARGC1A, G6PD, GHR, GH1, NEFH, EGR1, RUNX2, BGLAP, NEFM, NKX2-5, KDM6A, ADNP, KMT2D, TXNRD2, STAR, NNT, MRAP, TBC1D24, TUBB1, TRAPPC9, ADAMTSL1, B3GLCT, THRA, DUOXA2, GNB1, SLC5A5, GATA6, MC2R, PRKAR1A, PDE4D, DUOX1, GNAS, SLC26A4, TTF2, SOX3, SLC20A1, EPHA5, PHPT1, TTF1, SLC26A7, ASAH1, DECR1, SEMA6A, ENO2, DUOXA1, CSF1R, NLRP2, CHD7, DNAJC17, CNR2, NLGN3, DNAJB1P1, ALB, EFNA5, HLA-A, ZBTB20, NRGN, SERPINA7, JAG1, THRB, PRL, TNF, TRH, PAX2, VDR, HHEX, NOS2, NBN, KSR1, NCOR1, PDCD6, LEP, DNAJB1, ACHE
-
Retroperitoneal Fibrosis
Wikipedia
"Idiopathic retroperitoneal fibrosis: a discussion of the etiology". J. Urol . 94 (4): 385–90. doi : 10.1016/s0022-5347(17)63635-8 . ... PMC 4926988 . PMID 26860343 . ^ van Bommel EF (July 2002). "Retroperitoneal fibrosis" .
-
Intraductal Papillary Mucinous Neoplasm
Wikipedia
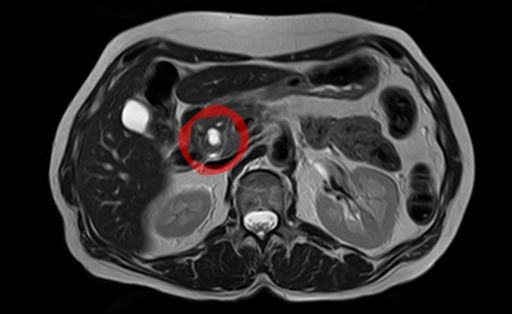
Diagnosis [ edit ] Histopathology of IPMN types in a distal pancreatectomy specimen from a 60-year-old man, by gross pathology (center image), microscopy and immunohistochemistry : The resected specimen (c) revealed that the mural nodule in the MPD consisted of PB-type IPMN with high-grade dysplasia (adenocarcinoma) (a) with a diffuse positivity of p53 immunostaining (an insert) and KRAS mutation (G12V). The BD-IPMN of the body was lined by gastric mucinous epithelium showing low papillary configuration with mild epithelial stratification with the same KRAS mutation (d), and the proliferation of similar gastric IPMN components sequentially involved the bottom of the mural nodule and the wall of the surrounding dilated MPD (indicated by red arrowheads) (b). The BD-IPMN of the tail was lined by flat, monolayer gastric mucinous epithelium lacking cellular atypia and KRAS mutation (e). [4] In most cases, IPMNs are diagnosed based on clinical and radiographic criteria. [5] If fluid from the cyst is aspirated, the CEA level is typically elevated. [5] Confirmation of the diagnosis with tissue is rarely necessary. [5] By histopathology , IPMN is characterized on light microscopy by Mucinous epithelial cells, [6] and growth within the pancreatic ducts . [7] Mucin 5AC is a useful immunohistochemistry marker. [8] Characteristic genetic alterations are those of KRAS and GNAS . [8] Further subtyping of IPMN can be done as either: [9] Gross pathology : Main duct, branch duct, and mixed duct lesions, which determines surgical management. ... A study using Surveillance, Epidemiology, and End Results Registry (SEER) data suggested that increased lymph node counts harvested during the surgery were associated with better survival in invasive IPMN patients. [15] Prognosis [ edit ] Survival 5 years after resection of an IPMN without malignancy is approximately 80%, 85% with malignancy but no lymph node spread and 0% with malignancy spreading to lymph nodes. [16] Epidemiology [ edit ] Side branch IPMNs are the most common pancreatic cysts. [5] IPMNs occur more often in men than women, and often occur in the 6th and 7th decade of life.GNAS, KRAS, MUC1, BRAF, ERBB2, SMARCA4, PIK3CA, ANXA10, LMO4, AIP, LONP1, GDF15, MSLN, POSTN, HHLA2, AQP4, SMUG1, VEGFA, SETD2, RMC1, NSD1, PPP1R2C, SNORD35B, SNORD14B, SNORD14C, SNORD14D, SNORD14E, MIA, ZEB1, TP53, MSH2, CFTR, CTNNB1, EGFR, FGF6, MSH6, IL1B, LEP, SMAD4, MUC2, TNF, MUC5AC, NPTX2, PIM1, PTEN, SNORD15A, SKIL, SPINK1, THBS2, MIR1290
-
Diabetes
Wikipedia
Of these two prediabetic states, the latter in particular is a major risk factor for progression to full-blown diabetes mellitus, as well as cardiovascular disease. [80] The American Diabetes Association (ADA) since 2003 uses a slightly different range for impaired fasting glucose of 5.6 to 6.9 mmol/L (100 to 125 mg/dL). [81] Glycated hemoglobin is better than fasting glucose for determining risks of cardiovascular disease and death from any cause. [82] Prevention [ edit ] See also: Prevention of type 2 diabetes There is no known preventive measure for type 1 diabetes. [2] Type 2 diabetes—which accounts for 85–90% of all cases worldwide—can often be prevented or delayed [83] by maintaining a normal body weight , engaging in physical activity, and eating a healthy diet. [2] Higher levels of physical activity (more than 90 minutes per day) reduce the risk of diabetes by 28%. [84] Dietary changes known to be effective in helping to prevent diabetes include maintaining a diet rich in whole grains and fiber , and choosing good fats, such as the polyunsaturated fats found in nuts, vegetable oils, and fish. [85] Limiting sugary beverages and eating less red meat and other sources of saturated fat can also help prevent diabetes. [85] Tobacco smoking is also associated with an increased risk of diabetes and its complications, so smoking cessation can be an important preventive measure as well. [86] The relationship between type 2 diabetes and the main modifiable risk factors (excess weight, unhealthy diet, physical inactivity and tobacco use) is similar in all regions of the world. ... Learning about the disease and actively participating in the treatment is important, since complications are far less common and less severe in people who have well-managed blood sugar levels. [88] [89] Per the American College of Physicians , the goal of treatment is an HbA 1C level of 7-8%. [90] Attention is also paid to other health problems that may accelerate the negative effects of diabetes. ... In addition, given the associated higher risks of cardiovascular disease, lifestyle modifications are recommended to control blood pressure. [93] [94] Weight loss can prevent progression from prediabetes to diabetes type 2 , decrease the risk of cardiovascular disease, or result in a partial remission in people with diabetes. [95] [96] No single dietary pattern is best for all people with diabetes. [97] Healthy dietary patterns, such as the Mediterranean diet , low-carbohydrate diet , or DASH diet , are often recommended, although evidence does not support one over the others. [95] [96] According to the ADA, "reducing overall carbohydrate intake for individuals with diabetes has demonstrated the most evidence for improving glycemia", and for individuals with type 2 diabetes who cannot meet the glycemic targets or where reducing anti-glycemic medications is a priority, low or very-low carbohydrate diets are a viable approach. [96] For overweight people with type 2 diabetes, any diet that achieves weight loss is effective. [97] [98] Implementing a plant-based diet may help manage or prevent type-2 diabetes by lowering glycated hemoglobin levels. [99] In addition, improvement in health may reduce the need for or number of medications needed by people who adopt a plant-based diet. [100] Medications [ edit ] Glucose control [ edit ] See also: Anti-diabetic medication Most medications used to treat diabetes act by lowering blood sugar levels through different mechanisms. ... Retrieved 25 March 2014 . ^ a b Kitabchi AE, Umpierrez GE, Miles JM, Fisher JN (July 2009). ... ISBN 978-1-5146-0305-5 . ^ a b Kitabchi AE, Umpierrez GE, Miles JM, Fisher JN (July 2009).INS, PPARG, CP, RAC1, APPL1, EIF2S3, FN1, ADIPOQ, SIRT1, SOD1, NR1I2, LEPR, IRS1, POMC, ATP6, PON1, CAT, PTGS2, AOC3, PAX6, IL2RA, CYBB, NR1I3, MAP3K5, MT2A, NCF1, MAFA, ALPK1, CPT1A, KIF1A, THBS2, NR0B2, ADRB1, GAD1, KIF5B, ZFP57, CLOCK, GLP1R, SERPINE1, PPARA, PTEN, ARNTL, ABCG2, ICA1, PROM1, LRP1, SLC22A12, KCNJ11, HNF4A, TCF7L2, UMOD, GLIS3, ABCC8, AQP4, AQP1, PTPN22, CDKN2A, STAT3, WFS1, HFE, IGF1R, CASR, APOE, F7, HNF1B, ASIP, GCKR, LMNA, IL6, PDX1, CFTR, LPA, INSR, IFIH1, PAX4, SLC19A2, HNF1A, FXN, HLA-DQB1, FTO, GCK, SRD5A2, HLA-DRB1, TP53, APOA5, HAMP, FOXP3, ATM, CDKN2B-AS1, COX2, CISD2, HLA-B, BRCA1, LIPC, BMP2, CAV1, GATA6, SPINK1, CUBN, PNPLA2, ARG1, CNBP, WRN, DNAJC3, GATA3, PRKAR1A, DNM1L, MLXIPL, GJA1, TRNL1, BLK, LIPE, TERT, PLIN1, IER3IP1, KRAS, ZFHX3, ITPR3, ELN, BSCL2, AIRE, SLC25A4, TLR1, SLC22A2, PDE4D, PRSS1, SLC30A3, DCAF17, NDUFS7, BRCA2, AGPAT2, C2, ND1, CPS1, HBB, ND5, SMAD4, PTF1A, MKKS, PRKAG2, HS6ST1, FOXC2, CYTB, FOS, SLC29A3, COX1, GNAS, FADS2, PIK3R1, MTNR1B, ND3, DKC1, CTLA4, TRNW, THSD7B, HPSE, PNPLA3, TRNV, HLA-DQA1, POLG2, ENPP1, NUBPL, HP, TXNIP, MAPK14, ACE2, TRNF, TRIM31, CYP2C19, RNASEH2B, HPN, SERPINF1, OR2B2, HLA-A, PALB2, PPARGC1A, CCDC28B, ND6, PARN, CCN2, SLC39A8, CTNNB1, TRNK, CDH23, TERC, HMOX1, EHMT1, HMGB1, TRNS1, ACE, CYP21A2, TREX1, USB1, ARMC5, DBP, CTC1, CAPN10, TRNQ, ELMO2, CST3, DAB2, NDUFAF5, DECR1, TRNS2, CARMIL1, SUGP1, RRM2B, NDUFA1, NDUFS6, MTOR, NEUROG3, PDE11A, TPRKB, NDUFS8, NDUFAF3, NDUFV2, FOXO1, ADIPOR1, NDUFAF1, NDP, NEUROD1, NFATC1, UCP1, FGF2, NFE2L2, UCP2, TIMMDC1, POLK, TTPA, TMEFF2, NGF, NHS, NOX4, KDSR, HIF1A, GLUL, TINF2, GIP, GJB3, NDUFB3, GH1, NDUFB9, ZBTB20, NDUFB10, NDUFS1, GLO1, GCGR, GCG, NDUFS2, NDUFS3, GATM, NDUFAF4, NDUFV1, NDUFA6, SAMHD1, CD274, GRHL1, GAD2, GABPA, NDUFS4, G6PD, FFAR1, RTEL1, PLA2G15, UCP3, MAML3, FOXRED1, ZCCHC8, CRP, NHP2, SIK3, KAZN, CELSR2, TRPV1, PALLD, TMEM126B, EDN1, OPA1, ACSS2, ANGPTL8, TNFRSF11B, CTRC, TWNK, RETN, DSPP, TGFB1, ATN1, SHROOM3, HGF, DPP4, SLC30A10, BEST1, NOP10, VEGFA, GPT, GPX1, FABP4, SEMA5B, NOS3, CCHCR1, NDUFB11, VDR, NPY, LEMD3, BCAS3, CDKAL1, TLR4, SARS2, ESR1, ROS1, TNF, WRAP53, RCBTB1, GSK3B, CHDH, ZFYVE26, EPO, GSTM1, OR12D3, PIK3CA, GPR101, APOA1, BBS1, GDF15, EIF2AK3, MIR146A, AVP, SLC5A2, ATP2B1, SLC5A1, SLC2A4, SLC2A2, SLC2A1, PTH, MIR21, ARSA, ST3GAL4, KL, SI, PTPN1, KCNQ1, APRT, APOC3, BBS2, BCHE, PRSS2, NSMCE2, PCSK9, PRKACA, WDR72, COPD, CAD, OR10A4, CAVIN1, MMP9, MAPK1, C9, MIR126, RAPGEF5, GOLGA6A, MMP2, MAPK8, NR3C2, ITPK1, LINC02694, BGLAP, BDNF, APOB, HMGA2, PRKAA2, SHBG, REN, ADRB3, RENBP, ZGLP1, HPN-AS1, PARP1, FADS1, CCL2, ALDH1A2, LPL, LRP6, ADAR, LIPC-AS1, ADA, LOC102723407, MC4R, PDE8B, ABO, ABCA1, IRS2, MBL2, AGER, LHX1, AGT, AKR1B1, MFAP1, PIP5K1B, USP8, AIP, BAZ1B, MOK, SELENBP1, MGAM, LCN2, CXCL12, AGTR1, ALDH2, ZPR1, LDLR, ALB, RBP4, LEP, AKT1, AHSG, LGALS3, PRKAB1, SLC34A1, IL1B, IL4, RNASEH2A, RBM45, SPP1, PLEKHH2, IAPP, LMNB2, IGFBP3, IL1A, COX3, PMM2, CPA1, ACCS, IL1RN, ND2, RNASEH2C, ARL6, POLD1, GJB4, FGF21, PIK3CB, PIK3CG, IDE, NLRP3, TRNC, UBE2Q2, PLCG1, IFNG, IGF1, PLG, NDUFA11, CETP, PIK3CD, NDUFAF2, MTHFR, XRCC4, UBXN11, ICAM1, POLG, FOXP2, SOD2, UBR1, CXCL8, IL17A, SLC30A8, PEX6, MPO, CASP3, PRKAA1, PEX1, PDILT, CERT1, HJV, IL10, IL18, CD34, CD36, HMGCR, ADAMTS13, CYP2E1, GHR, CYP3A4, EGFR, P2RY12, VWF, FGF23, TAC1, ALOX15, IFNA1, PDHX, SST, GPBAR1, P2RX7, IFNA13, HSPD1, DIANPH, TCF7, IGF2BP2, HSD11B1, SOST, SOCS3, RNF19A, ANGPT1, CALCA, KLF11, BCL2, SREBF1, UTS2, ADM, HPGDS, GSR, AHSA1, CRK, AIMP2, INSM2, GRAP2, SIRT3, ATP6AP2, MALAT1, HHEX, POLDIP2, TRBV20OR9-2, ACHE, AFP, DMPK, APP, FABP2, S100A8, APOL1, GAST, GFAP, S100B, REG1A, TNFSF10, MSTN, APLN, ADH1B, TRPM2, SELE, VCAM1, PLA2G7, G6PC2, CCK, FFAR4, PPARD, MME, TLR2, GSTT1, CD59, DDIT3, MIR155, TNFRSF1B, F3, IGFBP2, PYY, AR, PTPN6, PLAT, LAD1, SGK1, HSPA14, PTPRN, ADIPOR2, IGF2, EGR1, RFX6, KLK3, CXCR4, FGF19, FOXA2, MAPT, IDO1, CXCL10, MAP6, HBA1, KDR, OGA, NTN1, AKR1A1, HSPA5, HDAC3, KNG1, XBP1, PADI4, VEGFC, GNB3, IL2, LTA, SOCS1, SOAT1, CEL, PDK4, C20orf181, SCD, DMD, FBL, F2, NOS2, SERPINA5, MIR375, MIR223, PTK2B, NFKB1, ACR, SIRT6, PRKCB, NQO1, SPARC, ANPEP, RUNX2, PLXNA2, ANGPT2, G6PC, CCR2, IL22, AOC2, CYP19A1, APOA4, MIR34A, CTSD, PAEP, MIR29A, JAK2, CUX1, CYBA, DHX40, PINK1, B2M, PCK2, ATF4, MIR27A, PSIP1, PTX3, PTPRN2, MIR132, PCK1, BTF3P11, SPX, CCR5, ENTPD1, NAMPT, CD68, PPARGC1B, JAZF1, IGFBP1, PRKCA, SREBF2, IFNL3, BTBD8, SORCS1, INPPL1, TRMT10A, HSPA4, CNR1, MAPK7, TSPO, GSTK1, CNDP1, ISG20, SORBS1, CPOX, PECAM1, HSP90AA1, HSPB1, BMP7, MET, SLC17A5, CCL5, ANGPTL2, SORT1, BECN1, CLEC16A, ACACB, MTCO2P12, AGRP, NUCB2, ADRB2, OLR1, ERBB2, TNFSF11, TFPI, NR3C1, SMUG1, PERCC1, ACTB, ANGPTL3, S100A9, PDE5A, DPT, TFRC, HAVCR1, THBS1, TIMP1, APOM, S100A1, NRG1, SFRP5, P4HB, AMBP, SIGLEC7, MMRN1, HCAR2, RN7SL263P, ACP1, VPS51, NPHS2, KHSRP, MAPK3, ABCB6, SMAD3, MMP14, SMAD2, NAT2, TNFRSF9, SNCA, RIPK1, PEA15, CALR, MMP3, DGCR2, CASP8, ANXA1, LINC01672, LRP2, PPIG, LGALS1, SLC6A2, AHR, AKT2, HSPB3, MIR204, KMT2D, DCAF1, PTPN2, SELP, ARTN, MGP, POR, KLK1, AGTR2, SLC9A3, OGT, JAG1, BAX, GRK2, BRS3, SUMO4, DEGS1, LOX, MMP1, MICA, CARTPT, ERVK-18, S100A12, ABCG1, NOS1AP, FZD4, CEACAM5, PON2, HK2, SELENOS, DKK1, SIRT2, ATF6, EDNRA, HCLS1, HCRT, TH, TRIB3, DNMT1, HLA-C, TM7SF2, TREH, MNX1, WDHD1, REG3A, CD38, FN3K, HMGA1, HHIP, NOX5, CYP11B2, TKT, GSTP1, CYP2C9, ANGPTL4, GGT1, GFER, NOX1, SETD2, GAPDH, GAP43, NR5A2, FLNA, GOT2, FOXM1, UCHL1, NPPC, TXN, TTR, TRPC6, ADA2, ACSL1, TRAF6, NOTCH1, TNFRSF1A, ERN1, NPPB, CYP2J2, PRKN, RORC, GPR119, IL33, CLU, STK11, CHGA, FBXO32, IARS1, CECR, GIPR, TRIM63, RPAIN, CREBBP, SLCO6A1, CSF2, CCN1, HSPB2, STAT5B, CSF3, AGFG1, TRIM13, TCF3, IL2RB, PPP1R3B, PLA2G2A, PKD1, VIM, CCL4, PPIA, NPPA, MYD88, PLA2G1B, CCN3, MANF, RYR2, TNNI3, TYK2, SPG7, PLTP, PPP1R3C, ZFP36, PIN1, ST8SIA4, SMS, MT1A, PPID, PSMD9, MMP8, PDR, SLC6A3, PTHLH, ZEB1, PC, MIP, SLC6A4, TEK, PCSK1, PDCD1, PROX1, TFAM, PRL, TFE3, SLC22A1, MEN1, TIMP4, SELENOP, SYP, PGF, PGM1, RAF1, SELL, PLA2G6, RARRES2, CXCL5, SERPINA1, CCL21, SLC2A3, OGG1, TIMP3, TIMP2, DDAH2, PSAT1, ELANE, CASP1, ARID4B, UBASH3A, TLR9, CCKAR, TET2, ETV3, SLC52A1, SLC35G1, SLC47A1, IMPACT, EPHB2, CBLL2, EPHB1, MEG3, ENO2, ELK3, IL23A, CAPN1, ISYNA1, CFB, MIR15B, CABIN1, MIR15A, DDAH1, MIR145, CD2AP, MIR143, BGN, CANX, GNAO1, DESI1, SGSM3, FLT1, GAL, CA2, GP6, CD86, NRG4, ATP2A2, CISD1, CYP2D6, CYP2B6, CXADR, CX3CR1, CTSS, CTSB, HDAC11, ZC3H12A, CSN2, PRRT2, MUC16, SLC2A10, CMKLR1, CNTF, CPE, COL9A3, COPA, MUL1, CYP7A1, CHRM3, TRPM6, EDNRB, CD40, CD40LG, SLC2A9, DYRK1A, CD44, ACKR3, CIP2A, C1QTNF5, IL21, CDK4, TRPV4, DLD, CDKN1C, CHI3L1, DCN, CELA3B, COMT, CFH, KIR2DS2, MIR27B, FST, FOXO6, ITGAM, IDDM7, MIR29B1, GGTLC4P, IFNAR1, MIR29B2, GGT2, KIR2DL2, TUBB4B, KCNMA1, GGTLC3, AMPD1, IL6R, GGTLC5P, CXADRP1, MIR146B, PPIF, IL15, SH2B3, ELMO1, KEAP1, ITGB2, BMS1, SOX13, ADORA1, ADRA2B, INSRR, PGR-AS1, PER3, CDC123, HRC, MIR200B, TNDM1, MIR210, ARR3, KAT2B, ACACA, LNPEP, MIR217, HLA-DPB1, HSPA1A, HSPA1B, HLA-DQB2, ADCY5, C2CD4A, GGTLC1, GPR151, H3P10, MIR486-1, RPS6, BPIFA2, PWAR1, ABCC11, WNT3A, DNER, MIR483, RPS6KB1, C1QTNF3, CCL20, DBA2, TMX2-CTNND1, SALL1, SRL, UCN3, STAT1, SSTR5, LOC102724197, STAT5A, TMEM18, SSTR4, CCL11, SLC26A9, TRIM21, MRGPRX3, ST2, NOXO1, MRGPRX4, SOX2, MBOAT4, SLC18A2, SMPD1, MIR24-1, MIR22, GPIHBP1, PWAR4, STX4, ENHO, TBPL2, C2CD4B, C1QTNF12, LINC01194, SLC11A1, FRMD3, MIR203A, MIR122, MIR130A, MIR140, SLC6A8, MIR200C, MIR149, MIR19A, MIR199A2, MIR199A1, SLC5A3, MRGPRX1, MIR25, MIR503, MIR93, LYPD4, SP1, MIR499A, MIR184, PLB1, CREBRF, GPR166P, VN1R17P, MIAT, MIR17HG, SFTPD, SLC16A11, SHC1, MIR30D, TMPRSS6, OXER1, MIR302A, CRTC2, ZBTB7C, SNAP25, SNAI1, TPCN2, GPRC6A, IL27, TP63, CIDEC, STXBP1, SCGN, LILRB1, TCFL5, WNT1, NFAT5, CXCR6, KHDRBS1, CCT2, KCNQ1OT1, PRDX4, HYOU1, NSA2, RACK1, NOD1, SEMA3A, SLC22A7, BTN2A1, KLF2, BACE1, PAMR1, UBE2I, POC1A, TXN2, PRPF6, NUP62, UTRN, PDAP1, VDAC1, DAPK2, LPAR3, VGF, PES1, LPIN1, YY1, YWHAZ, STXBP3, F2RL3, TAM, S1PR2, GPR55, MSC, SLC33A1, LPAR2, WASF1, CD163, PLA2G10, SQSTM1, ARHGEF7, DENR, CES2, TNFRSF6B, KLF4, SLC7A5, RIDA, TNDM, TSHZ1, ZMYM2, AIR, CELA3A, ALMS1, NR1H4, HDAC4, GLP2R, BCAR1, NPEPPS, NR4A3, TBPL1, AIM2, COX5A, SH2B1, FETUB, FOXD3, IGAN1, EBF2, COP1, ABCG8, NOD2, IDDM17, SRR, BACH2, CORO7, LGR6, TFF3, UBL5, JPH3, TG, CAMK1D, SLC52A2, TCF21, INTU, TXNDC5, UBASH3B, SULT1A1, PPP1R15B, CCDC8, ANGPTL6, MFRP, SETD7, VTCN1, COL18A1, TACR1, TAPBP, TAT, TBP, LIN28A, TGM2, FAM20C, TRPV5, TWIST1, GHRL, UFM1, TSC2, TTN, DTL, TLR7, CYB5R4, ZNF395, DCTN4, SDF4, ASCC1, IL20, TBK1, IL17B, NR2C2, TPO, TPD52, ERRFI1, TREM2, KRT20, DLL4, TRPM7, ANO1, TMSB4X, TSPAN8, TSPAN7, TLR3, SOX6, STAP2, KDM3A, PAG1, SYN2, H3P40, HLA-DQA2, CELA1, NFKBIA, NGFR, NIDDM2, EIF5A, C3, NNMT, NOTCH3, EPOR, NPHS1, SLC11A2, NRF1, HDAC2, KLF9, BTC, NFKB2, PTTG1IP, NEFL, ECE1, CALCR, MYH9, CAPN2, MYC, MVD, DUSP1, CASP9, CASQ1, PRMT1, CAV3, MSX2, MST1, MSMB, BSG, OGN, HSF1, GYPB, PCNT, PCSK2, PCYT1A, F2R, GYPE, F2RL1, ATF3, ERBB3, GYPA, GUSB, GTF2H1, ARNT, F8, PGC, ATP2B2, EZH2, ATP5PF, PBX1, ATR, GZMB, PAWR, P2RY1, BCL2A1, EREG, HAS2, P2RX3, ERCC2, BMP4, BMP6, ERBB4, BMPR2, CD3E, CD14, SLC25A3, LAMP2, CTNND1, KRT16, COL1A1, CTSL, CYP1A1, LAMC2, CYP8B1, LIF, LBP, LCK, IL13, LETM1, CYP27B1, CXCR1, COL11A2, KRT5, COMP, KLF6, KISS1, IMPDH2, COX8A, CPB1, CSTA, KCNJ1, IRAK1, JUND, JUN, CSE1L, CRYZ, CRH, CRMP1, CD55, LIG4, CD19, DIO2, MDM2, CDK5, MECP2, DHPS, MEF2A, MEFV, MIF, LRP5, CD69, CD80, MS4A1, DPYD, HSPA9, DPYS, CDKN2B, DEFB1, IGFALS, MCL1, MC3R, DDT, IGFBP7, CEBPB, MB, MAS1, MAOA, MAFD2, CETN1, CTSC, CHAT, IL3, LYZ, PHB, ITPR1, FBN1, PMP22, FGFR4, GPER1, FASN, PON3, GPR35, FDPS, ADH1C, AMD1P2, ALAD, PVR, ADORA2A, AMH, RNASE3, RAD1, FAP, GCLC, PVT1, PROS1, GATA4, FKBP5, ADCYAP1, ALCAM, GLRX, FLG, FOXO3, ADD1, FLII, FMO3, GC, FKBP4, PSPN, ALOX12, ALOX5, GCH1, PROC, ALOX5AP, PSMA6, AMD1, PLAG1, ANXA2, GJA3, PTGS1, APOA2, ADRA2A, ADRA1A, GRN, F10, APLNR, PSPH, ADORA2B, FAS, ANXA6, GK, F11, CBLIF, FES, GPR42, FABP1, XIAP, GAS5, MFSD1, TFB1M, PROK2, GORASP1, PDIA2, WDR13, FGFR1, DAPK1, NIF3L1, ELOVL5, DIO3, THADA, GNPNAT1, SLC25A19, DLAT, AGXT2, DGKQ, FGF13, SCPEP1, DIH1, FBRS, DAPK3, DBH, TSPYL2, DES, DMRTA1, LHPP, TNMD, PCIF1, WNK1, DEFA3, DEFA1, DIAPH1, UBE2O, FGL1, STRA6, INSIG2, FHL1, ABCG5, DDOST, FHL2, PRDM16, DCX, RMDN1, METTL9, IKZF5, RTN4R, MOV10L1, UBE2Z, CSK, CERS2, CSH2, PDGFD, NRBP1, FLAD1, CDK5RAP3, SCUBE1, CTAA1, CYP4F12, SLC37A4, SH3KBP1, MOGAT2, ASRGL1, ERVW-1, CTBS, BICC1, COASY, CSH1, PADI1, CSF2RA, NME7, SEC61A1, MED25, CRY1, GPR132, MAP1LC3B, TSC22D4, CS, SLC38A1, KCNH6, NENF, CSF1, DSE, ZNF436, CSF1R, NT5C, PIK3R4, FUT2, CYP3A5, CYP4A11, F11R, CYP24A1, FOLR1, FOLH1, PAGR1, CYP27A1, FGF1, FSD1, MBOAT7, FLT4, DAB1, TMED7, TVP23B, ASPSCR1, AK3, NAA16, CTF1, CYP2D7, CTNS, NR5A1, FSHMD1A, VASH2, FOSB, SHCBP1, CTRB1, CTSG, MBL3P, CTSK, FOLR2, NLRX1, CYC1, ASAP1, CYP1A2, DLX3, CLEC1B, GER, SF3B6, ENG, MPC1, ZFR, ENO1, ENPEP, EP300, EMCN, HEMGN, FAT1, GIMAP5, FBXW7, RHOT1, ERCC1, RMDN3, MSTO1, RASD1, ELAVL2, SARDH, PLXNA3, EIF4E, ENAH, EIF4EBP1, SYBU, EIF4EBP2, FERMT1, ITLN1, GDE1, RNPC3, PIAS4, MIOX, TRIM33, NBAS, ATP6V1H, SERPINB1, MTPAP, FBLN1, MAP3K20, ERG, FABP3, FBLIM1, FEV, ARL15, H2BS1, EPB41L4B, TREM1, DDIT4, F5, ROBO4, SIAE, APBB1IP, SMOX, YIPF1, F9, CNNM2, FAM3B, SLCO1C1, CASZ1, ESD, MOCOS, MARCHF1, TUG1, ESRRA, ETS1, IL17D, DYM, NCAPG2, VPS13C, FABP5, HDL3, EXT2, IL20RA, FBN2, ELOVL3, EIF2S1, AKR1B10, SEMA6A, AHRR, AS3MT, DACT1, GJD2, DSC3, ADAMTSL3, PELI1, GOPC, RCAN1, KMT5AP1, DSG3, GPC4, PCBP4, DCDC2, CFAP97, FGB, NT5C3A, NEUROD4, DMRT1, DNAH8, ARHGAP22, OPRPN, GLRX5, PCTP, ZNF410, MZB1, DNASE1, DNM2, DNMT3A, MARK4, DOCK3, RNF213, COQ9, FCGRT, EGF, HBEGF, NSFL1C, EBF1, LANCL2, SIRT7, ZC4H2, EDA, S1PR1, GPRC5C, GSDMB, EEF2, ACOT13, USE1, EFNB2, EFEMP1, RAB14, MYDGF, PCDHGA4, ZNF253, SUCNR1, ADAMTS9, CHPT1, ENY2, NMUR2, SPHK2, DUSP4, DUSP9, E2F1, HSD17B7, FBP1, CLDND1, ASAH2, TOR1A, CYP26B1, RP9, LTB4R, SESN2, MIR200A, MIR214, MIR211, ARRB2, STS, MIR205, SERPINC1, ATIC, RERE, ATP12A, AREG, FXYD2, ATP2A1, MIR190A, MIR185, MIR182, MIR181C, ATP4A, MIR152, MIR216A, MIR221, RNASEK, MIR30A, MIR99A, MIR98, MIR96, ANG, MIR320A, MIR31, MIR30E, ANXA5, MIR29C, MIR222, APC, APCS, MIR296, APOH, AQP7, MIR23B, MIR23A, AQP9, MIR150, AVPR1A, MIR148A, GPR142, TMEM189, DST, LINC-PINT, KLF5, BUB1, TICAM2, SLCO4C1, ZACN, C1QBP, AVPR1B, SOX2-OT, CYCSP51, NANOS3, C4A, VWA2, TSPAN33, GADL1, ARMH1, TMEM189-UBE2V1, SERPINA13P, LINC01550, BNIP3L, AVPR2, BAAT, MIR142, BAD, MIR134, CCND1, HCN2, BCR, MIR107, MIRLET7G, GTF2H5, BID, SMIM10L2A, HES3, C1QL3, LIN28B, CCL4L1, AMY2A, CDNF, TMEM119, DNM3OS, ADAM10, TRPM2-AS, EMSLR, LINC01150, OPN1MW3, ADCY3, ARAP1-AS1, ADCY8, ERVK-20, ACVR2B, KLRC4-KLRK1, RBM14-RBM4, VIM-AS1, OCLN, ARAP1-AS2, PLIN2, MIR466, AAA1, MIR6835, ACP3, MIR330, LOC110806263, SERPINA3, H3P42, H3P28, H3P8, ABL2, LOC113664106, LINC02605, LOC111674464, ERVK-32, LOC102724334, ACADSB, ASIC1, CST12P, CERNA3, THRA1/BTR, ACP5, TP53COR1, CBSL, TMED7-TICAM2, C4B_2, H3C9P, MIR433, ALDH1A1, HCC, DELYQ11, ABCD2, NPS, MIR497, MIR409, HNP1, MIR449A, IBD20, MIR429, MIR424, MIR377, ALPI, ALPP, TRIM72, PLF, MIR338, ECSCR, SMIM10L2B, ALAS1, SFTPA1, KIF28P, SOD2-OT1, HOTAIR, TLE5, MIR675, PSS, AFM, AGA, OPN1MW2, POTEF, AIC, INS-IGF2, MIR618, MIR615, MIR579, H3P37, AIF1, BRINP3, C4B, CRISPLD2, AZIN2, CISH, MYSM1, CYGB, CKM, ERCC8, TXNRD3, CLCN7, TMEM54, SLC46A1, CHIT1, IL17F, CLK2, TP53INP1, CLPS, CCR1, ACKR2, GPSM2, SLC38A5, CHRNA4, C1QTNF7, OSCAR, THEM4, CD52, FOPNL, DEGS2, CYP2R1, MSS51, RLN3, CEBPA, DCD, TADA1, MARCHF3, ARAP1, LRG1, SNAP47, CENPA, ABHD15, RMI2, SLC5A11, GPR146, CMM, NAF1, MTG1, SLC41A2, KISS1R, ZGPAT, HOPX, SYVN1, CPT2, NLRC5, ASCC2, CR2, CREB1, CNP, TKTL2, MAGT1, ATF2, MIXL1, FSD1L, CREM, SETDB2, KCNK16, SPZ1, CLDN4, CPD, CPB2, CTBP1-DT, ARRDC4, CNR2, TIMD4, RFT1, COL6A3, PYGO2, BMF, KNSTRN, UCN2, GALP, KRT90P, CORT, PLXDC2, ORAI1, HAVCR2, ATP5MD, MSI2, CDKN3, C5AR1, FOLH1B, FNDC5, CAPN3, CAPS, CAST, CARS1, HOXA11-AS, PHACTR1, RANP1, LRRC55, CNIH2, NRK, PRSS55, CBLB, CBS, PAOX, ARID2, PTCRA, NLRP6, ANGPTL5, CALM3, TSACC, KSR2, MMAB, BPIFA4P, CA1, CACNA1A, RSPO1, ZNF763, CACNA1E, CHAMP1, HECTD4, TCERG1L, DDR1, NEAT1, SLC25A20, HS6ST3, CALB1, NEGR1, CALM1, CALM2, CCKBR, CD1D, SGMS2, UPRT, SERPINA12, PIWIL4, CACUL1, SIRPA, HT, ADGRE5, CDC42, CDH1, FUNDC1, CD5L, KLF14, TAAR1, WDR36, CDH5, CDH11, CDH13, CDK2, CDKN1B, CD47, SCARB1, WIPF2, CD33, CD8A, MARCHF10, SPRED1, LINC00599, AMOT, SLC2A12, PPM1K, KLB, UPP2, RMDN2, CD27, ERFE, CD28, SIK1, PDIK1L, ZNF569, CILP2, EEF2K, GAPDHS, NPC1L1, MTNR1A, WNT6, WNT5A, WNT2, MTTP, NSD2, MUC1, LAT2, MUSK, MMUT, VPREB1, MUTYH, VIP, EZR, VHL, MYF6, VEGFB, MYL2, PPP1R12A, NAB2, WNT7A, MTM1, MPI, XBP1P1, SCG2, FZD5, MPST, PAX8, TUBA1A, MPZ, PXDN, MRE11, ABCC1, MS, ZNF236, MSH3, SF1, MT1E, YES1, MT1JP, XRCC1, XPC, XK, VASP, KDM6A, NAGLU, USF1, NNAT, CRISP2, NM, TPM4, NME1, NOS1, NOTCH2, TNFAIP3, NRTN, YBX1, TLR5, NTS, NTSR1, NUMA1, OAS3, TLE3, TLE1, OAT, TJP1, NINJ1, TRPC3, NIDDM1, NCL, UPK2, UPP1, NUBP1, UGCG, NBN, NCAM1, NCK1, UCN, UBE2V1, TSC1, NDN, NFATC3, TNFSF4, NFATC4, NFIL3, TTC4, TST, TSHR, SLMAP, MPG, SLC16A4, LTBR, CCN5, NRP1, DPM1, TRIM24, SUCLA2, TNFRSF10B, TNFRSF11A, SIGLEC5, FADD, MXD1, SMAD1, TNFSF14, HRK, SMAD7, DGAT1, MAP2, VAMP4, DYNLL1, MBNL1, CCN4, LRPAP1, DDX39B, ASAP2, KRT18, SLC7A7, GPRC5A, KTN1, LAMP1, RPL14, CH25H, RPSA, TRPA1, LCN1, LCP1, EIF2S2, LDHB, FUBP1, SPHK1, LGALS2, LGALS3BP, LGALS4, LOXL2, NCOA1, ABCB11, MBP, MC2R, UBL4A, MGAT2, KITLG, AKAP1, HBHR, CXCL9, AAAS, MITF, MLH1, ECB2, MAP3K9, MAP3K10, STAM, MLN, FZD3, FOXO4, NTT, MMP10, MNAT1, TKTL1, AXIN1, BRAP, CST7, STC2, SMCP, MADD, ADAM11, DNAJB9, MAPKAPK5, LGR5, MDH2, ELP1, MFGE8, IRS4, MDM4, CUL1, MEHMO, EEA1, TAGLN2, MEOX2, LOH19CR1, ODC1, ODF1, ORM1, PSEN1, SLC5A5, PSMD7, PSMD10, SLC3A1, PTBP1, PTCH1, PTGDS, PTGIS, SLC1A3, SKP2, SIX1, PMEL, ST6GAL1, PTH1R, PTPRD, PTPRF, SGCA, SFTPC, SFTPB, PSEN2, PRSS8, TIE1, SLC7A2, POU4F1, SNRNP70, PPP1R3A, PTPA, PPY, PREP, PRKCD, SMO, SMARCA4, MAP2K3, MAP2K6, SLC20A2, SLC20A1, SLC19A1, SLC14A1, SLC12A3, SLC10A2, MASP1, SLC9A1, SRSF6, SRSF5, NECTIN1, SEMA3F, SATB1, RHEB, ACSM3, RHO, RIT2, S100A10, RNASE2, BRD2, RNY1, RYR3, RNY3, RXRG, RREB1, RPS19, ROBO1, RPS6KA1, ROCK1, RPL36A, RPL29, RHD, CLEC11A, SCN2A, CCL22, SELPLG, RAC2, RAG2, RAN, RAPSN, SDC4, RBP3, CXCL11, RCN2, SCN7A, OPN1LW, CCL16, REG1B, RELA, REST, RFC2, RGS1, SCN10A, POU2F2, POU2F1, SOD3, ADAM17, PAX2, TDO2, TDGF1P3, TRD, PCBD1, TCF19, PCDH8, PCMT1, PCNA, TCF4, TBXA2R, PDB1, TBX1, PDC, PDE7A, TAP2, TAP1, TALDO1, TAGLN, TERF1, PAPPA, PAM, TGFBR2, KLF10, THRB, THRA, THBS4, CLDN11, OXA1L, THBD, OXTR, TGFBR1, PRDX1, TGFBI, TGFB3, TGFB2, P2RX4, TGFA, P2RX5, PA2G4, FURIN, PDGFRA, PER1, SORD, SYT5, PLCB3, ITPRID2, PLEK, SRY, SERPINF2, PLK1, AKR1D1, PLN, SPRR2A, SPR, PLXNA1, SPINT1, PNOC, PODXL, SOX9, SOX4, POU1F1, SOS1, SORL1, PLAUR, PLAGL1, ST13, SERPINB6, SYT1, PF4, A2M, VAMP2, SUV39H1, PF4V1, ABCB1, STX1A, STIM1, ST14, ELOVL4, STC1, PIM1, PKLR, STAT4, PKM, STAT2, PKNOX1, MTA1, SLC16A3, ICOS, NFASC, UNC13A, H1-2, PDCD11, TPX2, HABP2, HARS1, MSRB2, HBG2, KLRK1, SACM1L, CARD8, INPP5F, NLRP1, ANKRD26, AAK1, VASH1, PUF60, MRAS, TUSC2, MYO16, KDM6B, NNT, PASK, GSN, GSTA4, BRD4, TRAM1, NPTXR, ABCB10, GTF2H4, TARDBP, SIRT4, ZNF629, CUX2, MCF2L, GYS1, PHLPP1, PLCB1, FBXO28, NBEAL2, TBC1D1, ATG4B, HCRTR1, ACOT7, PHB2, STK38, ESM1, SLC29A2, RAPGEF4, LIAS, KIF2C, SLC27A2, IMMT, METAP2, HNRNPK, EBNA1BP2, ERP29, PDIA5, BTG3, PRSS21, RALBP1, SUB1, PAPOLA, HOXA3, CPSF4, ONECUT1, NUP42, RPP14, AKAP13, RBPJL, PARK7, SLCO2B1, KLF12, UBE2K, DUSP12, PTGDR2, HLA-DMA, IRAK3, HNF4G, MAP4K5, SLC2A6, HLA-DMB, HMGCS2, SLC7A9, NR4A1, FAF1, PTPRT, CXCL1, RBFOX2, ARHGEF1, RPS6KA6, PDCD4, GPR162, HCAR1, GDF2, GLS2, NAAA, GDF10, CACYBP, TAF5L, FOXP1, SND1, LAT, GDNF, GEM, ATP2C1, GFPT1, GHRH, GHSR, GREM1, OPN1MW, GBE1, GRM5, MAT2B, TRPM5, GALNS, GALNT3, GAS1, DROSHA, CNIH4, STXBP6, GAS6, DBNL, FLVCR1, OSTM1, LAMTOR2, DEXI, REM1, TRBV7-8, DLL1, IGHD1-7, SLCO1B3, TOR2A, RPL10, GJA4, ANKRD2, GJA5, TPSG1, ARIH1, TAFA5, GCA, ZNF318, PANX1, TMEM245, RAB38, GPR39, FFAR2, GPX3, SH3BP1, GPX4, GRIA2, ELP5, GRIN1, GRIN2A, GRIN2B, GRM2, BACE2, MCHR1, PART1, GORASP2, FBXO8, FBXO25, PHGDH, GLA, LRIT1, GLI2, ACOT11, TKFC, SNED1, UTS2R, CHMP2B, TIPARP, KANK2, TPGS2, GLUD2, GNAT1, GP1BA, CXCR3, MMP24, MALT1, HOXC4, ING2, INSL3, RASSF2, INSM1, HDAC9, SART3, SEMA3E, IRF1, LPIN2, SOCS5, IKBKE, IRF7, ISL1, PRDX6, ITGAX, ITGB7, ITIH1, H6PD, CCL4L2, CIR1, PIEZO1, IP6K1, MRPS30, GIT2, PARP2, CHAF1A, INSL5, IL15RA, BCL2L11, HDAC6, CASP8AP2, FOXK2, CCS, ILF3, ILK, TNFSF15, IMPDH1, MVP, MFN2, RNF10, NUAK1, TBC1D4, ING1, EI24, BAG3, IVD, JAK1, LONP1, KCNJ9, SLIT2, KCNN4, B4GALT5, KIR2DL3, KIR3DL1, KLKB1, PIWIL1, STK17B, MFHAS1, NOG, PTTG1, HACD1, KRT8, ARHGEF2, PCSK7, HGS, PDCD5, KCNJ6, KCNJ5, KCNJ3, FHL5, GAL3ST1, ADAMTS1, ADAMTS4, MAPK8IP1, NAPSA, ROCK2, CHST3, PCYT1B, HOMER1, CIAO1, MAP4K4, JUNB, NCR1, KCNC2, CYP7B1, KCNE1, KCNH2, CD101, MAMLD1, GJC1, ABCC5, IDDM3, HSPG2, NDST1, HTC2, GNLY, HTR2A, CCT4, HTR2C, SLC35A1, TNC, IBD2, UBD, IBSP, ID1, DEAF1, CIB2, FBLN5, SEMA4D, CARM1, CAP1, SLCO1B1, SH2B2, IVNS1ABP, TRAF3IP2, USP19, HPX, HRH1, PGRMC1, WASF3, SEPTIN9, NEK6, DHS, GIPC1, HSP90AB1, MASP2, HES1, HSD3B1, HSD11B2, TNFSF13B, CELF1, HSPA2, POSTN, TIMM44, TOMM40, IL12B, VAV3, IL1R1, RABEPK, MRPS31, GDF11, DDX39A, NUTF2, IL4R, WASF2, IL6ST, MBNL2, IL7, IL7R, LRPPRC, RASGRP1, HIPK3, IL9, PTPRU, IL18BP, NR1H3, ABCC4, CDK2AP2, STUB1, IFNAR2, C1D, LYPLA1, RBM14, IDDM4, RAPGEF3, CPQ, PEMT, IDDM11, CORO2B, SIGMAR1, IGF2R, CITED2, MICU1, IGFBP5, TFG, LAMC3, TNIP1, LANCL1, A1BG
-
Chronic Bacterial Prostatitis
Wikipedia
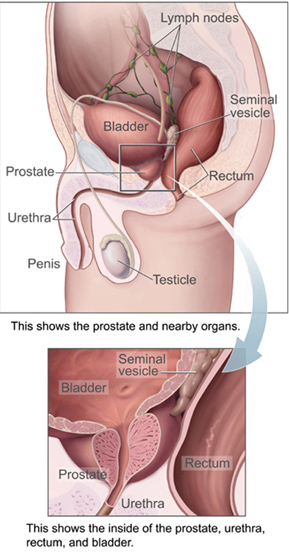
This is particularly true for gram-positive infections. [ citation needed ] In a review of multiple studies, levofloxacin was found to reach prostatic fluid concentrations 5.5 times higher than ciprofloxacin, indicating a greater ability to penetrate the prostate. [9] Clinical success rates with oral antibiotics can reach 70% to 90% at 6 months, although trials comparing them with placebo or no treatment do not exist. [10] Persistent infections may be helped in 80% of patients by the use of alpha blockers ( tamsulosin , alfuzosin ), or long term low dose antibiotic therapy. [11] Recurrent infections may be caused by inefficient urination (benign prostatic hypertrophy, neurogenic bladder), prostatic stones or a structural abnormality that acts as a reservoir for infection. ... A 2007 study showed that repeated combination pharmacological therapy with antibacterial agents (ciprofloxacin/azithromycin), alpha-blockers (alfuzosin) and Serenoa repens extracts may eradicate infection in 83.9% of patients with clinical remission extending throughout a follow-up period of 30 months for 94% of these patients. [17] A 2014 study of 210 patients randomized into two treatment groups found that recurrence occurred within 2 months in 27.6% of the group using antibiotics alone (prulifloxacin 600 mg), but in only 7.8% of the group taking prulifloxacin in combination with Serenoa repens extract, Lactobacillus Sporogens and Arbutin. [18] Large prostatic stones was shown to be related with the presence of bacteria, [19] a higher urinary symptoms and pain score, a higher IL-1β and IL-8 concentration in seminal plasma, a greater prostatic inflammation and a lower response to antibiotic treatment. [20] Additional images [ edit ] Prostate, urethra, and seminal vesicles. ... FEMS Immunology and Medical Microbiology . 60 (2): 99–112. doi : 10.1111/j.1574-695X.2010.00723.x . PMID 20698884 . ^ Nickel JC, Downey J, Feliciano AE, Hennenfent B (September 1999). "Repetitive prostatic massage therapy for chronic refractory prostatitis: the Philippine experience". ... "Clinical and Biochemical Influence of Prostatic Stones". Urologia Internationalis . 98 (4): 449–455. doi : 10.1159/000455161 .
-
Autoimmune Disease
Omim
The 4 affected persons shared an HLA haplotype: A1, C-, B8, DR3 and Dw3. Lippman et al. (1982) found a high frequency of autoimmune manifestations, both clinical and laboratory, in relatives of a proband with autoimmune hemolytic anemia, 1 with immune thrombocytopenic purpura and 8 with systemic lupus erythematosus (SLE; 152700).PTPN22, TPO, TG, IL17A, IL6, CD40LG, IFNG, IL4, SIRT1, FAS, CD28, STAT3, TNFRSF1A, TNFAIP3, FCGR2B, CXCR3, AICDA, SIAE, C4B, TNFRSF4, TIMP1, HP, PECAM1, TGFB2, B2M, PF4, FBL, EGR1, HMGCR, RBP3, F8, COPA, HPX, KRT19, RAPSN, SOD2, CRYAA, HSPA9, CRYBB2, COL1A1, ITGB6, C1S, HARS1, AHNAK, CCR6, TYK2, ICOS, CD226, IL23R, STAT4, CD40, IL2RA, IRF5, TSSK1B, TNFSF15, PTPN2, CD247, BACH2, UBASH3A, ERAP2, JAK1, IL2RB, ETS1, TNIP1, CAMK4, TRAF1, SH2B3, TAGAP, CARD9, IFI16, UBE2L3, ZMIZ1, CCR3, INS-IGF2, KIF5A, KIAA1109, REL, FLT3, ANKRD55, SMAD3, SH2D2A, FUT2, RASGRP1, VAV3, ATG16L1, ATG5, TNFRSF11B, NTRK1, KCNH7, SLC22A5, ICOSLG, AFF3, ATXN2, IL1A, SUOX, CLEC16A, SLC11A1, CCDC88B, PSMD5, IL17D, IL1B, LURAP1L, MB21D2, CRP, FAM107A, MPO, IL7, STING1, FOXP3, GEM, MYD88, MOG, ELMO1, CXCL10, IL27, GPR35, INS, FCGR3B, PRL, C1QTNF6, FCGR3A, FCGR2A, FAP, PUS10, TNFSF13, IL2, CIITA, IL1RN, SPATA13, NFKB1, IL23A, SLC30A10, IL18, CSF2, MMEL1, NLRP3, IL17F, ATN1, IL16, IGF2-AS, DAG1, IL15, GAD2, LINC02656, C12orf42, LINC02649, DPP4, GAD1, IL12B, CCN4, DDX6, ARID5B, RO60, TRIM21, HSPA14, ATXN2L, CXCL12, IL10, MIR146A, TNFSF10, IL37, MIR155, CTLA4, TLR7, ACKR3, CRB1, IL7R, FNBP1, ERAP1, STAT1, SOAT1, PLXNC1, IL9, TREX1, MBL2, SLC30A8, SPRED2, SPP1, C1orf141, YDJC, MIR3681HG, FASLG, AQP4, PTCSC2, LINC00824, LINC01250, LEP, TNFRSF13B, LINC01934, IL33, ANKRD30A, KLRC4-KLRK1, BCL2, NOD2, HMGB1, SAMHD1, BLK, CD274, HLA-G, CPT1C, PDCD1, PADI4, LPP, TTC34, BTK, NONOP2, ITGAM, TP53, VDR, IL12A-AS1, PIK3CA, PIK3CB, LINC00993, TNFSF13B, NKD1, HSPD1, KIR3DL1, LINC02605, IL21, ADCY7, PLEK, LRRK2, PLEKHA1, TNFSF4, RAB5C, RMI2, PHF19, AHR, TSHR, KRT20, PIK3CG, LINC02752, LINC02357, ALDH2, LINC02341, PIK3CD, MACIR, TMEM131, NEURL4, AIRE, LINC00271, IFIH1, FCRL3, IFNA1, HLA-C, HLA-B, HLA-A, CCL2, IFNA13, CD1D, NLRP1, TGM2, IRF1-AS1, CD19, MS4A1, CD22, CD86, HT, CD38, ADGRL2, TGFB1, CUTALP, SH2D1A, LRBA, TLR9, HAVCR1, DLEU1, IL22, HAVCR2, TRBV20OR9-2, RAB5B, KLRK1, MYDGF, ISG20, HLA-DQB1, MALT1, MROH3P, TNF, RBM45, ARHGAP31, CGAS, HLA-DQA1, TNFRSF1B, TLR4, HLA-DRB1, PRTN3, ABCB1, PRDM1, CR2, NUDT11, CYBB, PTPRC, ESR1, HSPA4, SUMO4, ZAP70, ENO1, JAK3, ADAR, HLA-DPB1, ARHGEF5, DSG3, KLK3, CBLB, TIMELESS, CD24, TLR2, MBP, S1PR1, TPMT, NCF1, LTA, SH2B2, RNPC3, EBI3, SYK, MMP9, WAS, IFNB1, APOH, ACE, ACAD8, STIM1, CDR3, IFN1@, HCRT, MTOR, NR3C1, SNCA, FCGRT, MAPK1, NFE2L2, VIP, LINC01193, CD44, TLR8, CYP27B1, CD47, DNASE1, IL18R1, MIF, CCR5, HNF1A, GRN, RUNX1, GPR42, GZMB, GLUL, CD80, PLAAT4, MIR21, CBLL2, SOCS1, ADIPOQ, VIM, HLA-DOA, PRKN, LPAR2, ADRA1A, ADRA2B, CXCR6, S100A9, LGALS9, RAG1, JAK2, SERPINA1, TNFRSF13C, LGI1, ANP32B, BRS3, VEGFA, HLA-DQA2, IDO1, IL34, MUL1, SLAMF1, CXCR4, DECR1, EDNRA, TNFRSF14, TNFRSF9, TAM, SSTR4, GFAP, PTEN, AFP, TBK1, DOCK8, BCL6, ITGB2, MRGPRX3, TPI1, C4A, MS, NR0B2, CCR2, C5AR1, GPRC6A, GIMAP5, CAMP, SLC22A4, TNFRSF12A, ICAM1, IRF8, S100A8, ADRB2, MFGE8, MRGPRX1, CNTNAP2, ICA1, FLI1, MRGPRX4, LGR6, AKT1, REG1A, HSPA1B, FZD4, APCS, AIM2, PSIP1, BTG3, IL13, CASR, MIR17HG, MBTPS1, CEACAM5, LINC01194, BANK1, CCL21, OXER1, CSK, PTX3, CCR7, LILRA3, COL17A1, ZFP36, TBC1D9, FCGR1A, ORAI1, TNFRSF8, DNASE1L3, MYO9B, LILRB4, MAVS, SSB, GABPA, PSMB8, GPR151, GPR166P, LPAR3, VN1R17P, IL24, KIR2DS1, ITGAL, KIR3DL2, LGALS1, INPP5D, ITGAX, CXCL8, IRAK1, IFNAR1, TNC, KLRB1, LGALS3, CLEC4A, CD69, PRKCQ, DDX58, TNFSF14, NM, MAP4K3, IL17B, MAPK14, EGR2, MIR142, IRAK4, CD55, DBP, HSP90AA1, MUC1, DSG1, NR4A2, ROBO3, TBX21, CALR, ACP1, AGER, HRH4, USO1, PPARG, LOC102724971, ANXA1, LOC102723407, PTPA, HPSE, SEC14L2, CEACAM1, ARID5A, NXF1, MERTK, MTHFR, DNMT1, GNAO1, NT5C1A, CFH, CD83, CCL3L1, BTLA, HLA-DQB2, GCK, MSC, HLA-DRB3, IL32, HMOX1, SELL, NR4A1, FMR1, PTPN6, FLNB, LGALS8, IL1F10, HSPA1A, ZC3H12A, FOXO1, PDCD1LG2, BTNL2, CD276, MEFV, THSD7A, FOXJ1, HIF1A, PML, TP63, KRAS, SPPL2A, PGF, ERVK-6, POMC, LSM2, MRC1, RNF19A, CD46, MST1, CD200, COX2, PSMB9, LAG3, TNFAIP8L2, NFATC2, PTGS2, NFAT5, FAM167A, NOS2, PELI1, SMAD7, NT5E, RAB4A, IKZF4, IGHV3-52, POU1F1, SIRPG, IFNL1, DNASE2, POLDIP2, LCE3C, LCE3B, GPR183, EGR3, DST, DEK, SLC20A1, MIF-AS1, MAP4K1, MICA, HDAC9, BCR, RPL17-C18orf32, IFNL2, ESR2, HSP90B1, ESRRB, CXCL13, COPD, EZH2, SPATA2, ENTPD1, C3AR1, LRPPRC, CASP1, TLR5, IL20, IL21R, CDKN2A, CD34, CD14, CD52, CTSC, CHI3L1, DEF6, CD3E, CCR4, MBL3P, ADAM17, COL4A3, MIR326, KLRG1, CRK, STAT6, STAT5B, CTNNB1, CYP21A2, ADAMTS13, SEMA3A, GDE1, F9, PADI2, RORC, SERPINB3, AHSA1, LRR1, TRAF3IP2, DEXI, AGT, MTCO2P12, PARP1, HRES1, RPL17, SLAMF6, TREM1, IGHG3, ADA, ACTB, TYRO3, IL6ST, SOCS3, TMEM39A, IRF1, DNER, RAG2, HLA-DRB4, HLA-E, ROCK2, GH1, TRAF6, ISG15, FLII, C9orf72, AIMP2, SGK1, TIGIT, AR, GUCY2C, NUDT10, IKZF1, ALB, CCL5, AMPH, GRAP2, TYR, TRAF5, TYRP1, KDM6A, TPM3, HSP90B2P, TTN, RGCC, SGSM3, VPREB1, TLR3, PRKAR1A, PREP, THM, VAV1, SUGP1, STIM2, SP100, TFF3, SLC7A5, CD244, RTEL1, POU2AF1, CXCL5, NR4A3, CCL20, SCO1, ATF7IP, OTUB1, PTPRN, PRKDC, PTPRN2, TET2, SAMD9, RPS19, NECTIN1, FEZF2, RGS1, NEIL3, NUDT15, SHBG, PTGER4, GSDMB, PMEL, TCF7, CNTN2, TAP2, TAP1, PROC, ST2, SSRP1, GPR65, LEF1, DPYSL5, SPG7, MANF, SH3BP4, PSMD7, FOXD3, SLC5A5, DTL, IL17RA, LST1, SELE, H3P28, GLIS3, DEFB1, EGFR, CELSR3, BCL2L11, ABCB6, TSC22D3, IL1RAPL1, DLAT, DHODH, OPTN, POLR3A, CD160, VGLL3, CNR2, G3BP1, CYP24A1, MIR125A, CYP2B6, CXADR, MIR150, CST3, MIR17, TRIM13, MIR204, MIR22, EIF4E, CELA1, ELANE, WDHD1, RC3H1, GRIA3, GPER1, GPI, KAT2A, GAS6, LRRC32, GAPDH, SIGIRR, ACKR1, USP18, SOX13, FOLR2, NCR3, EXT1, ETV5, ERN1, ERBB3, ERBB2, EPO, EPHB2, NR1I3, ARMH1, CORT, MIR223, SIRPA, ARRB2, C5, TEC, FST, SERPING1, BSG, C4B_2, LILRB1, CXCR5, BAK1, LINC-ROR, ASCL2, ARR3, CNGB1, APOA1, BIRC3, ANXA5, AIF1, AHCY, JAG1, ERVK-32, ADAM10, ACP3, H3P13, MASP2, CXADRP1, CALCA, MIR499A, CASP3, CMKLR1, MIR23B, MIR29B1, MIR29B2, RABEPK, PRSS16, CDKN2D, LILRB2, LANCL1, CD74, CD72, CD70, CD59, TNFSF8, CD5L, CD4, CD1C, SERPINH1, WG, TRIM22, CAV1, EBNA1BP2, NOD1, CXCL1, IGHE, NR1I2, NRAS, RPSA, TNFSF12, NFKBIL1, MARCKSL1, LAIR1, LAD1, NOS1, KRT14, NOS3, KDR, KCNA3, OTULIN, JUNB, BMF, HARS2, ITGA4, PNP, MPZ, TIMD4, NEFM, LIF, GPR174, MDM2, ABCC1, WLS, COTL1, HDAC11, TARDBP, TNFRSF6B, CD84, MECP2, CFLAR, LPL, RIPK2, LYZ, HDAC3, LRP4, SYVN1, VTCN1, TNFSF9, LRP1, BCL11B, IRF7, TNFRSF25, HSPE1, IRF3, VSIR, SMOC2, PROK2, HRH1, IL1RL1, HOXC6, PLA2R1, ARHGEF2, HNRNPA1, MORC3, HNF4A, PKM, PLCG2, HLA-DRB5, PDLIM7, CD200R1, PAEP, NR3C2, IL22RA2, CLEC7A, PDX1, DEPTOR, ORMDL3, SEMA5A, IL6R, IL5, IL3, ARHGEF28, IL2RG, IGF1, IDDM8, SMUG1, TNPO3, CCR9, BRD4, UTS2, HCST, IGHV3-69-1, CLCF1, PDE10A, KLRA1P, IGHV3OR16-7, CD300C, SNAPIN, IFI44L, TSSK2, CARD8, TNRC6A, SLC7A9, ARHGAP26, PHGDH, HECW1, KIF21B, CLEC4E, MON2, ERAL1, PKP3, HRH3, SP140, DUSP12, FGF21, WDTC1, HSPB8, VSIG4, IL36RN, HBP1, STK38, MMRN1, ARL5A, CDC42EP1, HPGDS, LAT, DICER1, GCA, TNFAIP8, PRDX5, JMJD6, SS18L1, TNFRSF21, IKZF3, DSTN, RIPK3, WWP2, ACSBG1, CNTRL, DUSP14, AGO2, CIC, TAB2, DKK3, CDK19, CABIN1, NLRC5, TRIB2, IFNL3, MLKL, NLRC3, CRTC2, PRSS55, OTUD1, JAZF1, ADGRG3, VAMAS6, YTHDF3, NPNT, SGMS1, MILR1, THEMIS, GSDMA, TREML4, CLEC4D, TSPAN33, SYCN, TGM6, IRGM, TICAM2, AIS1, GSTK1, MALAT1, PGAM5, IDO2, GIMAP7, IFNLR1, EGLN2, PRRT2, TIRAP, LYPD1, GRIN3A, CYP2R1, RHEBL1, H4-16, PLD4, SOCS4, ZPBP2, SPNS2, NLRP5, TNFAIP8L1, IL17RE, SLCO6A1, DEFB104A, BPIFA2, SMCR8, PWAR1, SIX5, PLB1, ANKFN1, IL31, ILDR2, CCDC22, MIR3614, H4C15, MIR20B, MIR146B, HNRNPA1P10, MIR584, KIR2DS2, HLP, DEFB4B, MIR1238, MIR1908, TMED7-TICAM2, RAB4B-EGLN2, RTL1, ERVK-20, PSC, ERVK-18, LINC01882, LOC105379528, GSTT1-AS1, RN7SL263P, SIRT1-AS, EEF1AKMT4-ECE2, H3P23, H3P24, DEFB104B, MIR424, MIR423, MIR382, CCL4L1, LOC390714, ST20, LINC00951, MIR140, MIR148A, MIR15A, MIR182, MIR183, MIR199A1, MIR199A2, MIR20A, MIR221, MIR30A, MIR320A, MIR34A, MIR93, MIR96, TNFSF12-TNFSF13, DEFB103A, MDD1, MIR340, CISD2, EPSTI1, FOXQ1, FOXP2, XKR8, PTOV1, IL20RA, IL20RB, DNAJC10, APBB1IP, EGLN1, BCOR, PXK, BANP, LAMTOR1, MARCHF1, RAVER2, BPIFB1, QRSL1, FBXW7, LGR4, PRO2268, GPRC5D, VAC14, ZNF415, PAG1, ASH1L, GPRC5C, DEFB103B, RTRAF, TRIM33, HDAC7, POMP, DCPS, FLVCR1, C1GALT1C1, IL19, EFEMP2, ERVW-1, CXXC1, MINK1, CD207, NOX3, DELEC1, F11R, TRAT1, TMED5, TMED7, GAL, CRYL1, CHCHD2, DCXR, MZB1, TFDP3, UBAP1, SLC25A37, SUCNR1, RETN, SPHK2, SLC52A2, RTL10, MANEA, CCDC134, KIAA0319L, NAA25, ARMC9, TET1, TSGA10, MPIG6B, HM13, UNC93B1, FCRL4, GPR61, ZCCHC7, HVCN1, CARD11, SPZ1, INSM2, FCRLA, TMEM60, TRIM5, SYS1, ZNF804A, NLRX1, FCRL2, C1GALT1, MMP28, DUSP22, PCBP4, CD248, CD177, SLURP1, ABHD6, GATAD2B, SERINC1, TAOK1, ZFAT, NCOA5, ZNF410, CXCL16, IGAN1, GAS5, BCORL1, PDLIM2, AZI2, IL25, CDCP1, PINK1, RAPH1, DHX58, YME1L1, ABCA4, POLD3, HGF, HLA-DRB2, HLA-DRA, HLA-DPA1, HLA-DOB, HLA-DMA, HK1, HELLS, HMGN1, HDAC1, GZMK, GYPA, GTF2B, GSTT1, GSTM1, HLA-F, HMGN2, IFIT3, HSPA1L, CFI, IDDM6, IAPP, HSPG2, HSPA8, HSPA2, HES1, FOXA1, PRMT1, HRH2, HPRT1, TLX1, HNRNPD, FOXA2, GSN, GRM3, GRIN2A, FOXO3, FOSB, FOS, FOLR1, FLT3LG, MLANA, FLG, FKBP1B, GRIN1, FKBP1A, FHL1, FH, FEN1, FCN2, FCN1, CENPI, FUCA1, FUCA2, FUS, FUT1, SLC37A4, GATA3, GC, GCG, GCH1, GFER, GFI1, CBLIF, GPC3, GPR25, GPT, GPX1, IFI27, IGFALS, NFATC1, MAS1, MAP3K1, MEF2D, MDK, MCM3, MCL1, MCAM, MARK1, MICB, MAN2A1, MAG, SMAD4, M6PR, LTBR, LTBP3, KITLG, CXCL9, IGFBP3, MYH1, NCF4, NCF2, NCAM1, NARS1, NAIP, MYH9, MUTYH, MLH1, MTX1, MTTP, ATP8, ATP6, MSH2, MMP1, LTB, LRP2, LPA, IL12RB1, ITK, ITGAE, IRF6, IRF2, ING2, ILK, IL9R, LMNA, IL4R, IL1R1, IKBKB, IGHA1, CCN1, IGFBP6, ITPA, ITPR3, JUN, JUND, KARS1, KIR2DL2, KIR3DS1, KLRC1, KLRD1, KNG1, KRT1, KRT10, KTN1, LAMB1, LBR, LGALS3BP, LIG4, FABP4, F5, F3, C2, CA5A, CA1, FMNL1, C4BPB, C4BPA, C3, C1QA, CACNA1A, C1QBP, TSPO, BTC, ZFP36L1, BRAF, BMP7, CA6, DDR1, F2R, CAV2, CD8A, CD3G, CD2, CCT6A, CBL, RUNX3, CAT, CALCR, CASQ2, CASQ1, CASP10, CASP8, CAPZB, CALML3, BMP5, BMP2, BMI1, AGRP, AMH, ABCD1, ALCAM, AKT2, NR0B1, AGTR1, ACAN, CFB, ADRB1, ADM, ACTA1, ACHE, ABR, ABO, ANPEP, ANXA6, BIRC2, APOE, AREG, ARG1, ARNTL, ARSD, ARSL, ATP4A, ATP5PO, AVP, AXL, BACH1, BAX, BCL3, BDNF, CD27, CD36, CDK1, DPT, DUSP2, DUSP1, DTX1, DSC2, DSC1, SLC26A3, DNMT3A, DDIT3, DIO2, DIAPH2, DHFR, DHCR7, GSDME, DEFB4A, E2F2, ECM1, EGF, ELAVL2, ELAVL4, ELK3, ELN, MARK2, ENG, ENO2, ENO3, EPHA4, EPHB1, NR2F6, ERG, ERV3-1, EXT2, DDX5, GADD45A, CDC25C, CLU, CREB1, CR1, COX8A, MAP3K8, COL9A3, CNR1, CHRNA4, DCT, CHI3L2, CFTR, CDR2, CDK9, CDK8, CDH1, CREBBP, CREM, CRYGD, CSF3, CSF3R, CSN2, CCN2, CTSB, CTSG, CTSV, CTSS, CYP2C19, CYP2D6, CYP2E1, CYP3A5, CYP19A1, DCC, NEU1, NFIL3, DCTN6, H4C1, H4C8, H4C3, H4C11, H4C12, H4C6, H4C4, EOMES, H4C5, H4C9, NCOA3, MADCAM1, AP3B2, KMT2D, FOSL1, H4C2, H4C13, EIF3C, KHSRP, RNASET2, RTCA, PDE8B, TNFSF11, PRKRA, PDLIM4, YARS1, H4C14, BHLHE40, IKBKG, PARG, CILP, PLA2G6, LOH19CR1, CSRP3, DDX39B, SCLC1, TSPAN7, TRP-AGG2-5, TRAF3, CRISP2, TPP2, TOP1, TNP1, TIAM1, IL1R2, THBS1, TH, TGFBR2, TGFBI, TGFB3, TERT, TRPC1, TRPM2, TSC2, TTR, TXN, TYROBP, UBE2D3, UBTF, USP4, VCAM1, WFS1, WNT5A, XRCC1, YWHAZ, ZBTB16, RNF112, DNALI1, KCNK5, RIPK1, NFKB2, BCAP31, DDX39A, PSME3, ATP6AP2, TRIM28, TOB1, NAMPT, CERT1, TRIB1, TSPAN32, PTPRU, IL18BP, HDAC6, DGCR2, GAB2, PSMD14, AKR1A1, TNFRSF18, ZNRD2, KLF1, CELF2, KHDRBS1, RBCK1, SLCO1B1, BATF, KAT5, CCL26, SEMA4D, IFI30, PRMT5, CPQ, KLF2, ABCA7, TOX, ECE2, SOCS5, KAT2B, F2RL3, MBD2, BCL10, EIF2B5, SPHK1, IER3, ALKBH1, RGS6, SOCS2, GMPS, NRP1, IL1RL2, IL18RAP, TNFRSF11A, ARTN, PSTPIP1, CBFA2T2, PDCD5, MYOM2, OSMR, LRRFIP1, STK17B, S1PR2, CD163, RPL23, ABCG2, GSTO1, IL27RA, GOSR1, CLOCK, APOBEC3B, TMBIM6, TDO2, TBP, PRKCA, MAP2K2, MAP2K1, MAPK3, PKN2, PRKCD, PRKCB, PRKAB1, PRLR, PRKAA2, PRKAA1, PRF1, PPP2R2B, PPL, PPARD, EIF2AK2, PROS1, TALDO1, PSME1, PTPN12, PTPN11, PTPN1, PTN, PTGER2, PTCH1, PSMD13, PRSS1, PSMD10, PSMD9, PSMD4, PSMB10, PSMA6, PSG1, PPARA, POU2F1, PON3, OCA2, SERPINE1, P4HB, P2RY11, P2RX7, OSM, SLC22A18, OAS1, PNN, YBX1, NPR3, NOTCH3, NME1, NGF, NFKBIA, PAK2, PAX4, PBX1, PDE3B, PDE4A, PDGFRA, VIT, SLC25A3, PIGA, PIM1, PIK3R1, PIN1, PLA2G1B, PLD1, PLG, PLP1, PMP22, PTPN13, PTPRM, PVR, SLC2A1, SIGLEC1, SMARCE1, SLC22A1, SLC6A8, SLC6A2, SLC3A2, SLC1A5, XCL1, SKI, SH3BP2, SRSF1, SELPLG, SELP, SDCBP, SNRPD1, SOD1, SOD3, SOS1, ABL1, SPIB, SPR, SPTAN1, SST, ST14, STAT5A, STK3, AURKA, STK11, STYX, TBXT, TACR3, SDC1, CCL24, NECTIN2, RBBP6, RIEG2, RHEB, RHD, RFX1, REN, OPN1LW, RASGRF1, CCL22, RASA1, RANBP2, RAF1, RAD51B, RAC2, RAC1, RNASE3, RORA, RPA2, RPS6KB2, RPS26, RRAS, S100A1, S100A10, S100A12, S100B, SAA1, SAA2, SAFB, SATB1, SCD, CCL17, CCL19, SPARC
-
Abortion In Australia
Wikipedia
Before then abortion law was for many years governed by case law under sections 82–84 of the Crimes Act 1900 of New South Wales . ... The offence is called "killing unborn child" and can be committed only around the time of childbirth [80] in Queensland , [81] Western Australia , [82] and the Northern Territory . [83] It is called "causing death of child before birth" in Tasmania . [84] In South Australia, it comes under the heading of " abortion ". [85] The definition is somewhat broader in the Australian Capital Territory , [80] [86] and comparably broad to English law in Tasmania [84] and South Australia . [85] [80] The offence was abolished in Victoria by the Abortion Law Reform Act 2008 (Victoria) . [87] New South Wales does not have a child destruction enactment, [80] but the Crimes Amendment (Grievous Bodily Harm) Act 2005 (NSW) amended the Crimes Act 1900 (NSW) so that s 4(1)(a) now defines "grievous bodily harm" as including "the destruction (other than in the course of a medical procedure) of the foetus of a pregnant woman, whether or not the woman suffers any other harm". [88] This was further amended by the Abortion Law Reform Act 2019 to "the destruction (other than in the course of a medical procedure or a termination of a pregnancy in accordance with the Abortion Law Reform Act 2019 ) of the foetus of a pregnant woman, whether or not the woman suffers any other harm." ... On the other hand, many women who have medical abortions performed at private hospitals may not claim the Medicare rebate. [89] South Australia is the only state which collects and publishes data on abortions. ... Less than 2% took place at or after 20 weeks. [89] Public opinion [ edit ] Main article: Societal attitudes towards abortion Since at least the 1980s, opinion polls have shown a majority of Australians support abortion rights, [90] and that support for abortion is increasing. [91] In 1987, a Saulwick poll found only about 7 per cent of Australians would not approve of abortions under any circumstances. [92] In 2003, a poll by The Australian Survey of Social Attitudes (AuSSA) found that 81% of Australians believe a woman should have the right to choose an abortion, and 9% believe they should not; 7% were neutral and 2% could not decide. [91] In 2005, a Nielsen Corporation poll found that 56% of Australians thought the abortion laws in place, which generally allow abortion for the sake of life, health, or economic factors, were "about right", 16% want changes in law to make abortion "more accessible" and 17% want changes to make it "less accessible". [93] In 2006, a poll by Roy Morgan Research found that 65% of the Australians approved of surgical abortion and 22% disapproved, and that 62% believed RU-486 should be available to women while 31% believed it should not. [94] In 2006, interviews found that 80% of Australians disapproved the use of sex-selective abortion . [95] In 2007, a poll by AuSSA found that 4% of Australians are opposed to abortion in all circumstances, 33% believe abortion should be allowed in certain circumstances and 57% believed it should be readily available whenever a woman wants one; 7% were undecided or did not respond. [96] In 2009, a study of polls conducted during Australia's 2007 federal elections found that a clear majority of both Labor Party and Liberal Party voters support abortion rights. [96] The study also showed that 77 per cent of winning candidates in the 2007 election favoured an unrestricted approach to abortion. [97] In 2010, a study published in The Medical Journal of Australia found that 61% of Australians said abortion should be lawful without question in the first trimester of pregnancy, while 26% said it should be lawful depending on the reason. [98] In the second trimester and third trimesters, support for outright lawful abortion was 12% and 6% respectively, while 57% and 42% respectively said it depended on the circumstances. [99] See also [ edit ] Abortion in New Zealand Abortion law Child destruction References [ edit ] ^ a b "Abortion Law in Australia" . ... Criminal Code Act Compilation Act 1913 (PDF) (v14-b0-05 ed.). 27 June 2009. p. 142 . Retrieved 31 March 2009 . ^ "Criminal Code Act – Notes" .
-
Dysphrenia
Wikipedia
References [ edit ] Chouinard G, Jones BD. Neuroleptic-induced supersensitivity psychosis: clinical and pharmacologic characteristics.
-
Facial Femoral Syndrome
Wikipedia
J Clin Ultrasound 42(1):49-52 ^ Daentl DL, Smith DW, Scott CI, Hall BD, Gooding CA (1975) Femoral hypoplasia--unusual facies syndrome. J. Pediat. 86: 107-111 External links [ edit ] Classification D OMIM : 134780 MeSH : C537916
-
Macroglobulinemia, Waldenstrom, Susceptibility To, 1
Omim
All persons with WM and all but 1 with autoimmune manifestations had HLA haplotype A2/B8/DRw3. A lod score of 4.86 favored linkage to HLA on chromosome 6p21 and a gene predisposing to lymphoproliferative and autoimmune disorders. ... Sanger sequencing identified MYD88 L265P in tumor samples from 49 of 54 patients with Waldenstrom macroglobulinemia and in 3 of 3 patients with non-IgM-secreting LPL (91% of all patients with LPL). MYD88 L265P was absent in paired normal-tissue samples from patients with Waldenstrom macroglobulinemia or non-IgM LPL and in B cells from healthy donors and was absent or rarely expressed in samples from patients with multiple myeloma, marginal-zone lymphoma, or IgM monoclonal gammopathy of unknown significance.MYD88, IL6, BTK, PAX5, CXCR4, BCL2, IGH, KRT20, CD19, MS4A1, MIR155, TP53, CXCL12, IRAK1, STOML2, IL4, IRF4, UCHL5, CD40LG, CD27, HAS1, MYOM2, USP14, SDC1, CDR3, SYK, TP73, ZAP70, IL10, ANPEP, MALT1, NAMPT, TNFSF13B, LPL, TNFSF13, MYC, PIK3CD, SMUG1, AKT1, BACH2, MTOR, AICDA, FCGR3A, FCER2, BCR, TCL1A, RAPH1, ARHGAP24, SLC28A1, EGLN3, TNFSF10, TNFRSF13C, TNFSF11, ARID1A, PRIMA1, IGHV3OR16-7, AIMP2, GRAP2, SLC35B2, LAPTM5, MIR206, MIR23B, XPO1, XBP1, VWF, VEGFA, MIR363, LOC102723407, CLEC12A, HDAC9, TCL1B, ANP32B, POLDIP2, IBTK, RNF19A, IGHV4-34, MAPK8IP2, TNFAIP3, ACSBG1, IGHV3-69-1, IGHV3-23, CD274, BLNK, ZHX2, ADAMTS13, IRAK4, LEF1, TLR7, AHSA1, COLEC10, CXCL13, EGLN1, EXOC2, TNFRSF13B, PXN, TNF, CD70, HCK, HAS2, GLI2, FGFR3, EFNB2, CTRL, CTNNB1, MAPK14, CRP, CRK, CD52, CDKN2A, CD79B, CD79A, CD40, HMMR, CD38, TNFRSF8, CD22, CD6, CCND3, CBL, SERPING1, BSG, BRAF, BLM, BCL9, BCL6, CCND1, ANXA5, HCLS1, IL1B, AURKA, PLCG2, STAT5B, STAT5A, SPIB, SPI1, SOAT1, CCL3, RAF1, PTPRC, MAPK3, MAPK1, PRKCD, POU2F2, POU2F1, POU2AF1, PIK3CG, IL2RA, PIK3CB, PIK3CA, NOTCH2, NOS2, NOS1, NCAM1, CD200, MMP8, MCL1, KIT, ITGAM, ITGA4, ISG20, IL4R, LOC102724971
-
Folate Deficiency
Wikipedia
Journal of the Royal Society of Medicine . 91 (2): 72–3. doi : 10.1177/014107689809100205 . ... Nutrition . 13 (11–12): 975–7. doi : 10.1016/S0899-9007(97)00340-7 . PMID 9433714 . ^ Kelly GS (June 1998). ... British Journal of Haematology . 113 (3): 579–89. doi : 10.1046/j.1365-2141.2001.02822.x . ... PMC 6814158 . PMID 31684687 . ^ Czeizel AE, Dudás I, Vereczkey A, Bánhidy F (2013). ... PMC 4695937 . PMID 26562127 . ^ Czeizel AE, Dudás I, Vereczkey A, Bánhidy F (2013).DHFR, MTHFR, CBS, MTR, BCL2, PCBP1, SLC46A1, MTRR, IL2, PAX3, TAGLN, NOTCH1, PCNA, MYOG, PSEN1, QDPR, RFC1, ROS1, SHMT1, MYC, SLC19A1, ADH5, TBX1, TCN2, TYMS, H3-4, SQSTM1, KHDRBS1, DKK1, FAM215A, NUP62, DCTN4, WLS, SERPINA13P, MIRLET7G, MIR34A, CBSL, TGFB1, MTHFD1, ALDH2, FHIT, APOE, BRCA1, BRCA2, C5, C5AR1, CRP, CSTB, CTSL, CYP2J2, DNMT1, DVL1, MARK2, ESR1, FOLR1, LPL, FPGS, GLI2, GNAS, GPT, GTF2H1, GZMM, H2AX, HSPA5, IFNG, IGF1, IGF2, IL10, LEP, RN7SL263P
-
Vaginal Bleeding
Wikipedia
Fertility and Sterility . 95 (7): 2204–2208.e3. doi : 10.1016/j.fertnstert.2011.03.079 . ... Fertility and Sterility . 95 (7): 2204–2208.e3. doi : 10.1016/j.fertnstert.2011.03.079 . ... American Journal of Obstetrics and Gynecology . 201 (1): 12.e1–12.e8. doi : 10.1016/j.ajog.2009.04.024 . ... "Challenges of diagnosing and managing the adolescent with heavy menstrual bleeding". Thrombosis Research . 143 : 91–100. doi : 10.1016/j.thromres.2016.05.001 . ... Journal of Midwifery & Women's Health . 54 (6): 483–91. doi : 10.1016/j.jmwh.2009.08.007 .
-
Chronic Obstructive Pulmonary Disease
Wikipedia
The diagnosis of COPD should be considered in anyone over the age of 35 to 40 who has shortness of breath , a chronic cough, sputum production, or frequent winter colds and a history of exposure to risk factors for the disease. [22] [27] Spirometry is then used to confirm the diagnosis. [22] [75] Screening those without symptoms is not recommended. [76] Spirometry [ edit ] Spirometry measures the amount of airflow obstruction present and is generally carried out after the use of a bronchodilator , a medication to open up the airways. [75] Two main components are measured to make the diagnosis, the forced expiratory volume in one second (FEV 1 ), which is the greatest volume of air that can be breathed out in the first second of a breath, and the forced vital capacity (FVC), which is the greatest volume of air that can be breathed out in a single large breath. [77] Normally, 75–80% of the FVC comes out in the first second [77] and a FEV 1 /FVC ratio less than 70% in someone with symptoms of COPD defines a person as having the disease. [75] Based on these measurements, spirometry would lead to over-diagnosis of COPD in the elderly. [75] The National Institute for Health and Care Excellence criteria additionally require a FEV 1 less than 80% of predicted. [27] People with COPD also exhibit a decrease in diffusing capacity of the lung for carbon monoxide (D LCO ) due to decreased surface area in the alveoli, as well as damage to the capillary bed. [78] Evidence for using spirometry among those without symptoms in an effort to diagnose the condition earlier is of uncertain effect, so currently is not recommended. [22] [75] A peak expiratory flow (the maximum speed of expiration), commonly used in asthma, is not sufficient for the diagnosis of COPD. [27] Severity [ edit ] MRC shortness of breath scale [27] Grade Activity affected 1 Only strenuous activity 2 Vigorous walking 3 With normal walking 4 After a few minutes of walking 5 With changing clothing GOLD grade [22] Severity FEV 1 % predicted Mild (GOLD 1) ≥80 Moderate (GOLD 2) 50–79 Severe (GOLD 3) 30–49 Very severe (GOLD 4) <30 A number of methods can determine how much COPD is affecting a given individual. [22] The modified British Medical Research Council questionnaire or the COPD assessment test (CAT) are simple questionnaires that may be used to determine the severity of symptoms. [22] Scores on CAT range from 0–40 with the higher the score, the more severe the disease. [79] Spirometry may help to determine the severity of airflow limitation. [22] This is typically based on the FEV 1 expressed as a percentage of the predicted "normal" for the person's age, gender, height, and weight. [22] Both the American and European guidelines recommend partly basing treatment recommendations on the FEV 1 . [75] The GOLD guidelines suggest dividing people into four categories based on symptoms assessment and airflow limitation. [22] Weight loss and muscle weakness, as well as the presence of other diseases, should also be taken into account. [22] Other tests [ edit ] A chest X-ray and complete blood count may be useful to exclude other conditions at the time of diagnosis. [80] Characteristic signs on X-ray are hyperinflated lungs, a flattened diaphragm , increased retrosternal airspace, and bullae , while it can help exclude other lung diseases, such as pneumonia , pulmonary edema , or a pneumothorax . [81] A high-resolution CT scan of the chest may show the distribution of emphysema throughout the lungs and can also be useful to exclude other lung diseases. [23] Unless surgery is planned, however, this rarely affects management. [23] A saber-sheath trachea deformity may also be present. [82] An analysis of arterial blood is used to determine the need for oxygen; this is recommended in those with an FEV 1 less than 35% predicted, those with a peripheral oxygen saturation less than 92%, and those with symptoms of congestive heart failure. [22] In areas of the world where alpha-1 antitrypsin deficiency is common, people with COPD (particularly those below the age of 45 and with emphysema affecting the lower parts of the lungs) should be considered for testing. [22] Chest X-ray demonstrating severe COPD: Note the small heart size in comparison to the lungs. ... Many people with COPD mistakenly think they have asthma. [36] The distinction between asthma and COPD is made on the basis of the symptoms, smoking history, and whether airflow limitation is reversible with bronchodilators at spirometry. [83] Tuberculosis may also present with a chronic cough and should be considered in locations where it is common. [22] Less common conditions that may present similarly include bronchopulmonary dysplasia and obliterative bronchiolitis . [80] Chronic bronchitis may occur with normal airflow and in this situation it is not classified as COPD. [23] Prevention [ edit ] Most cases of COPD are potentially preventable through decreasing exposure to smoke and improving air quality. [17] Annual influenza vaccinations in those with COPD reduce exacerbations, hospitalizations and death. [84] [85] Pneumococcal vaccination may also be beneficial. [84] Eating a diet high in beta-carotene may help but taking supplements does not seem to. [86] A review of an oral Haemophilus influenzae vaccine found 1.6 exacerbations per year as opposed to a baseline of 2.1 in those with COPD. [87] This small reduction was not deemed significant. [87] Smoking cessation [ edit ] Keeping people from starting smoking is a key aspect of preventing COPD. [88] The policies of governments, public health agencies, and antismoking organizations can reduce smoking rates by discouraging people from starting and encouraging people to stop smoking. [89] Smoking bans in public areas and places of work are important measures to decrease exposure to secondhand smoke, and while many places have instituted bans, more are recommended. [17] In those who smoke, stopping smoking is the only measure shown to slow down the worsening of COPD. [90] [91] Even at a late stage of the disease, it can reduce the rate of worsening lung function and delay the onset of disability and death. [92] Often, several attempts are required before long-term abstinence is achieved. [89] Attempts over 5 years lead to success in nearly 40% of people. [93] Some smokers can achieve long-term smoking cessation through willpower alone. Smoking, however, is highly addictive, [94] and many smokers need further support. The chance of quitting is improved with social support, engagement in a smoking cessation program, and the use of medications such as nicotine replacement therapy , bupropion , or varenicline . [89] [91] [93] Combining smoking-cessation medication with behavioral therapy is more than twice as likely to be effective in helping people with COPD stop smoking, compared with behavioral therapy alone. [95] Occupational health [ edit ] A number of measures have been taken to reduce the likelihood that workers in at-risk industries—such as coal mining, construction, and stonemasonry—will develop COPD. [17] Examples of these measures include the creation of public policy, [17] education of workers and management about the risks, promoting smoking cessation, checking workers for early signs of COPD, use of respirators , and dust control. [96] [97] Effective dust control can be achieved by improving ventilation, using water sprays and by using mining techniques that minimize dust generation. [98] If a worker develops COPD, further lung damage can be reduced by avoiding ongoing dust exposure, for example by changing their work role. [99] Air pollution [ edit ] Both indoor and outdoor air quality can be improved, which may prevent COPD or slow the worsening of existing disease. [17] This may be achieved by public policy efforts, cultural changes, and personal involvement. [62] A number of developed countries have successfully improved outdoor air quality through regulations. ... Excessive oxygen; however, can result in increased CO 2 levels and a decreased level of consciousness. [168] Corticosteroids by mouth improve the chance of recovery and decrease the overall duration of symptoms. [2] [62] They work equally well as intravenous steroids but appear to have fewer side effects. [169] Five days of steroids work as well as ten or fourteen. [170] In those with a severe exacerbation, antibiotics improve outcomes. [171] A number of different antibiotics may be used including amoxicillin , doxycycline and azithromycin ; whether one is better than the others is unclear. [84] The FDA recommends against the use of fluoroquinolones when other options are available due to higher risks of serious side effects. [172] There is no clear evidence for those with less severe cases. [171] For people with type 2 respiratory failure (acutely raised CO 2 levels) non-invasive positive pressure ventilation decreases the probability of death or the need of intensive care admission. [2] Additionally, theophylline may have a role in those who do not respond to other measures. [2] Fewer than 20% of exacerbations require hospital admission. [62] In those without acidosis from respiratory failure, home care ("hospital at home") may be able to help avoid some admissions. [62] Prognosis [ edit ] Chronic obstructive pulmonary disease deaths per million persons in 2012 9–63 64–80 81–95 96–116 117–152 153–189 190–235 236–290 291–375 376–1089 Disability-adjusted life years lost to chronic obstructive pulmonary disease per 100,000 inhabitants in 2004. [173] no data ≤110 110–220 220–330 330–440 440–550 550–660 660–770 770–880 880–990 990–1100 1100–1350 ≥1350 COPD usually gets gradually worse over time and can ultimately result in death.MMP1, SFTPD, HMOX1, HDAC2, TGFB1, MMP9, SERPINA1, FAM13A, DSP, TNF, EPHX1, MTCL1, CXCL8, EEFSEC, IL6, ELN, NOS3, SOD3, CYP1A1, NOS2, FOXO3, CXCL1, HTR2A, TRPV4, CYP1A2, RAPGEF3, TP53, TNNT2, MMP14, KLF5, CXCL2, CD8A, VEGFA, TIMP1, MUC5AC, IL17A, MIR218-2, NFE2L2, EGFR, CASP3, SERPINE1, PLAU, MUC1, SLPI, STAT4, MPO, CCN2, SMAD4, DDIT3, BNIP3, IREB2, FAIM2, CHRNA5, CHRNA3, CFTR, SMPD3, CASP8, CASP12, ERBB3, PLAUR, FAS, FASLG, NQO1, AGER, CHRNB4, MMP3, CTLA4, HSPA4, CHRM3, SCGB1A1, PLB1, PRTN3, THSD4, CYBA, RNF150, HSPA1A, NFKB1, HTR4, TNS1, PDZD2, WIPF1, PSORS1C1, HSPA1B, CDH13, HLA-DPB1, HLA-DPA1, DNAAF4, GSTCD, FTO, HSPA1L, ARHGEF38, INPP5D, GAB2, HYKK, DOCK1, CSMD1, NPNT, KCNK1, INTS12, CYBB, DNAH5, DLG2, TAFA2, RARB, GRIK4, CYS1, CFAP221, RAB4B, COPD, RSRC1, CASC15, PPARG, FAM227B, GABPA, CRACR2B, POFUT1, GP1BB, PRKCB, ZFPM2, HNF1A-AS1, GM2A, LAMA1, GC, NNT, C1orf185, NWD1, TSHZ3, NIFK-AS1, GSTT1, MIR99AHG, MKS1, GTF2I, ZBTB38, ATXN7L1, HIRA, GSTP1, GSTM1, BMP8A, SYCP2L, EFCAB5, MFHAS1, PTGS2, LINC01118, HLA-B, TGFB2, JMJD1C, PKD2L1, MMS22L, DELEC1, CFH, PKN2, FGL1, NRG1, HIF1A, MICAL3, ZKSCAN1, GLA, SCFD2, PLCE1, EML4, PCDH9, BNC2, LRMDA, ADGRV1, TNRC6A, ARMC2, ZDHHC18, HPGDS, MFAP2, ABLIM2, RBMS3, PCM1, TBX1, TOX2, PDE4A, TET2, FEV, IL33, LPO, MMP12, TSPAN14, HSBP1, SLMAP, CYBC1, HOOK2, SPAG16, HMGA2, PCDH15, SYNPO2L, DENND2D, STN1, TMEM254, NCF4, NCF2, PPP1R2C, CEP70, MYH7, SYN3, ADAM33, CASZ1, RSPH6A, BCAS3, PLXNA4, WDR20, KCNG4, ADAM19, IL13, PIK3CD, IL10, CDC123, NOL4L, PITPNA, ARHGAP42, IL5, ADIPOQ, IL1B, IL1A, LYSMD4, SLC35F3, CNTN4, IFNG, HERC1, AMZ1, SPPL2C, GLIS3, SCLT1, UBR3, TMEM219, ITPK1, PTPN22, CDRT15P1, TNPO1, SGF29, KCNMA1, CCDC69, GDAP1, BTBD9, ITGB8, IL18, KLHL32, GALNT13, SERPINE2, ITGA1, BTBD11, PIK3CA, HHIP, CXCL10, SNTG2, OR6F1, FGD6, MAPT-AS1, DIRC3, LINC00886, BTBD1, CAT, ZDHHC20P2, SLC30A10, ASTN2, CD96, C1GALT1, AMD1P3, ACE, WTAPP1, TBX4, HDAC7, BTC, TESK2, VGLL4, RREB1, SGCD, PPIEL, USP24, TLR2, CRP, SUZ12P1, LINC02869, IRF1-AS1, MAPK14, TM9SF4, CSE1L, PLF, ATP2C2, SFTPB, CCL2, TLR4, CDH11, LINC00598, SEC24C, NCF1, CCL28, FBXL7, LINC01006, CFDP1, COMT, ELANE, LINC01807, ANXA11, ANXA5, CHRM3-AS2, LINC01876, ADCY5, ADAMTSL3, LINC02863, COX10-AS1, RNASEH2A, LINC01937, LINC02625, ALB, RNPEP, LINC01997, ADRB2, HSF2BP, TOP2B, RNF220, CCDC91, PDSS2, MIA-RAB4B, RERE, DMWD, THRA, RAB4B-EGLN2, ARVCF, DPP6, WAS, NNT-AS1, CDYL, UFD1, MIR4527HG, SIRT1, ADGRG6, PIK3CG, MIR146A, STAT3, PIK3CB, MAPK1, TSLP, CHI3L1, IL1RN, SLCO6A1, EDN1, GSTK1, ICAM1, AHSA1, CDKN2A, MMP2, POLDIP2, SERPINA3, RNF19A, AIMP2, GRAP2, CRK, MYDGF, MIR21, IL22, CSF2, FOXP3, IL4, IL27, VIM, GSTM2, CTNNB1, VIP, MSTN, SPP1, CXCR2, MAPK3, ACTB, CD40, SOD2, NR3C1, HMGB1, GDF15, SOD1, CAMP, SLC6A4, IFNB1, MUC5B, SLC26A4, COX2, TIMP2, LEP, MTCO2P12, MAFD2, IFNA13, IFNA1, MIR145, CCR5, BCL2, ITGAM, VDR, IL17D, CYP2A6, MBL2, EGLN2, CTSS, DEFB4A, POSTN, NR1I2, SMAD2, GCLC, DNER, PPARGC1A, BCHE, NLRP3, CFLAR, SMAD3, NOTCH1, XRCC5, LTB, PTAFR, ADO, CHIT1, MIR34A, PTPA, PTX3, EGR1, MIR223, H3P10, CCL5, FGF7, AHR, SLC27A5, FOXO1, TLR3, PARP9, GLP1R, FN1, PTEN, PARP1, CST3, DEFB4B, AQP5, CEL, DPP4, LCN2, IL17C, PLAAT4, MRC1, FGF10, MCAM, PINK1, DEFB1, TAC1, ABCC1, CCND1, MARCO, LINC01672, NR3C2, ANPEP, CD163, ANXA1, TLR9, GCG, BTBD8, MMP7, PDCD1, EGF, BDNF, MOK, EPAS1, ITGB2, YWHAZ, CCL11, PIM3, CD68, SFTPA1, ABCB1, IL5RA, IL26, IL2, PWAR1, SLC52A2, SQSTM1, GZMB, ROBO3, PLA2G1B, CD274, FGF23, ADIPOR1, CHRNA4, TUG1, CSF3, CLCA1, CCR6, CX3CL1, TFAM, SOCS3, HP, CXCL5, ENTPD1, SAA2, MMRN1, NPPB, IL17F, KDR, NLRP1, CXCR3, F2R, KL, ABCA1, SLC2A1, JAK3, SIGLEC14, DDX58, GDF11, FBXO32, STAR, CASP4, PIGR, S100A8, S100A9, SAA1, CCR2, CD14, CD80, CD207, STAT6, SRC, P2RX7, RELA, P2RY2, SORD, HSPA14, SOX5, WNK1, PACC1, WNT5A, NAT2, CCL18, SLC6A2, IL25, SDHB, CIP2A, XRCC1, PI3, TRPV1, MAPK8, SFTPC, CCL1, IFIH1, BPIFA1, PON1, PPARA, APOM, BEST1, CCL22, CCL20, CCL13, PGF, SAGE1, PDGFRB, PDE7A, SOAT1, SNAI1, PRKN, SEMA6A, UGCG, PTPN1, RORA, PTGS1, PTH, DCTN4, IMPACT, GORASP1, PDE4B, CLEC7A, MEG3, RNASE3, HOXA6, TNFRSF11B, CYP2E1, CYP1B1, CTSG, CTH, TBC1D9, VCAN, CS, MIR29B1, MIR29B2, CPOX, COX8A, HDAC9, TIMP4, COL4A3, POTEKP, CHRNA1, CYP2B6, CYP3A5, F2RL1, MIR206, MIR126, MIR132, COX5A, EPHB1, MARK2, ELK3, MIR150, MIR155, HBEGF, SARDH, DMD, AIM2, FHL5, MIR191, DBP, HDAC5, S1PR5, POTEM, UCA1, CXCL13, ERICD, ALK, ALDH2, AKT1, AGT, ADRB1, ADM, KHDRBS1, ADAM8, ACTG2, ACTG1, CXCR6, SDS, ABCA4, APCS, SIRT6, ATF3, CAV1, MIR542, MIR570, CD34, CD86, SFTPA2, CD1C, CASP1, ATF4, CALCA, ZGLP1, SERPING1, ACOT7, GPR182, WDHD1, OGG1, MIR125A, FABP5, MBTPS1, WNT4, KRAS, KNG1, ITGAL, ITGA5, HDAC3, GSTO2, IRF3, IRAK1, INSRR, IL12B, IL11, PART1, BAMBI, MIR106B, PLA2G2D, LGALS3, HOXA13, BECN1, EHMT1, NOS1, NM, SYT1, MYOG, MYC, MUC2, COX1, MSR1, SESN2, TRIM63, MCL1, SMAD7, SIGLEC9, SH2D1A, HSPD1, TNC, HOXA11, MTOR, SMUG1, GPR15, GCLM, SUMF1, GALNS, MSC, ACTBL2, BRD4, GTF2H1, SERPINA13P, FLNA, FKBP5, FGFR4, CD83, FENDRR, FGF2, NUP62, GSR, HOXA1, HOXA@, HOXA10, HOXA9, HOXA7, HOXA5, HOXA4, HOXA3, SRF, HLA-A, HGF, CH25H, HCK, HBE1, PRKAG2, COMMD10, TLR7, TLR8, ATP6V1D, TRPV2, A2M, IRAK4, P2RX2, HEY1, RHOQ, SIRT3, TSPYL4, CYFIP1, CUL9, SARM1, PPRC1, MYCBP2, PHLPP2, SIRT2, SAR1B, KLRK1, MLXIP, TUSC2, PDAP1, PARK7, NUP42, ADAMTS13, WDR5, ABHD2, PIM2, FAM215A, PPP1R15A, PLA2G15, IL17RA, ASCC1, HEBP1, NTM, ASAP1, NOX4, DUOX2, RHOD, MYLIP, TBK1, SETD2, KLF15, DLL1, MAT2B, SIGLEC8, IL37, CYFIP2, INTS1, SH2B1, TPSG1, DCDC2, SPDEF, GULP1, YTHDC1, ISYNA1, MIR195, MIR149, MIR15A, MIR15B, MIR181A2, MIR181C, MIR186, MIR190A, MIR192, MIR197, MIR133B, MIR20A, MIR203A, MIR210, MIR212, MIR214, MIR22, MIR29C, MIR34C, MIR141, MIR134, MIR122, MIR10A, FOLH1B, ZEB1-AS1, MARCHF8, NT5DC1, IFNL2, IFNL3, IFNL1, H19, STPG4, CELIAC2, GADL1, CFAP100, SNHG5, SNHG17, PLEKHM3, MIRLET7C, MIR106A, LINC01555, MIR338, PGAM5, ELFN1-AS1, LINC00987, MIR3620, SMIM31, LINC01082, KLRC4-KLRK1, P2RX5-TAX1BP3, MIR4709, LINC-ROR, RORA-AS1, MIR374A, ADAMTS9-AS1, LOC102724334, CBSL, HLA-DQB1-AS1, ORI6, DINOL, LINC02605, COPDA1, PCNA-AS1, MIR2054, MIR664A, MIR1307, MIR380, MIR422A, MIR424, MIR146B, MIR495, MIR499A, MIR503, MIR483, MIR455, MIR552, MIR637, POTEF, MIR675, CDKN2B-AS1, MIR942, SOD2-OT1, LINC00861, CRYGEP, ADAMTS19, IL23A, MIB1, KLHL7, AGPAT3, SDHAF3, CAMK1D, PELI1, ARFGEF3, NLN, MAVS, CFAP97, TRPM8, PPP4R4, HAMP, ZNF410, CXCL16, SCAF1, IL21, SRR, NOD2, ZC4H2, ACSS2, CENPJ, ZNF692, GHRL, DUOX1, H2BS1, TREM2, TREM1, PID1, MOCOS, ATG16L1, SLC52A1, ANO1, ELP3, MTPAP, YY1AP1, HJURP, MAML3, SOX6, KLK15, DEPTOR, MMP28, AMOTL1, UCN3, RSAD2, BPIFB1, ORAI3, MUC16, ORMDL3, ARHGAP12, AZIN2, TXNRD3, MIR155HG, FSD1, SLC22A12, CD200R1, IL31RA, FUNDC1, ROMO1, LAYN, IL23R, DAB2IP, ESAM, BMF, WNT3A, PNPT1, PAGR1, ADIPOR2, NLRX1, HAND2-AS1, MAP3K19, DNAJC5, SPX, SETD7, HCG4B, FBXO38, CCNL2, FSD1L, MAGT1, DOT1L, AFAP1L2, ACCS, IL17RC, PAPOLA, SP1, MRPS30, FBL, GPER1, GPX1, GPX3, GPX4, GRIK2, CXCL3, GSK3B, GSN, GSS, GYPA, GYPB, GYPE, GZMA, HAL, HAS2, HCRT, HDLBP, CFHR1, HLA-DRB3, HMGA1, NR4A1, FOXA2, HNRNPD, HSD11B1, HSPA2, HSPB1, HSPB2, GOT2, GLRX, GLB1, FLT1, ETS2, MECOM, F10, FBLN1, FCGR1A, FDPS, FES, FGF13, FGFR1, FHL1, FKBP4, FOXC1, FOLH1, GHSR, FOS, FOSB, FPR1, FPR2, FSHMD1A, GAST, FUT8, GAD1, GALC, GATA4, GDNF, GH1, HTR2B, HYAL1, IAPP, LGALS9, KCNK3, KIT, KLK1, KLKB1, KRT5, KRT7, KRT18, KRT19, KRT34, LAMP1, LBP, LEPR, LIF, JUND, LMNB1, LOX, LRP5, LTA, LTC4S, TACSTD2, SMAD6, MAP1B, MAS1, CD46, SMCP, MDK, KCNJ2, JUNB, IDS, IL10RA, IFIT1, IGF1, IGFBP3, IGFBP7, CCN1, IGHE, IL1RAP, IL2RA, IL6R, IL7, CXCR1, IL9, IL12A, JUN, IL13RA1, IL15, IL15RA, IL16, TNFRSF9, IRF7, ISG20, ITGA2B, ITGAV, ITGB3, ITGB5, JAK1, ESR2, ERCC1, FGL2, EPO, OPN1SW, BICD1, BID, BMP4, BMP6, BMPR1A, BMPR2, BPI, BSG, BTF3P11, TSPO, C3AR1, C4A, C4BPB, C5AR1, CA2, SLC25A20, CAD, CALCR, CALR, CBS, CCNE1, CD1A, CD1D, CD5L, CD28, TNFRSF8, BAX, AZU1, AZGP1, ALPP, ACHE, ACLY, ACP3, ADA, ADCY2, ADORA2A, ADORA2B, AEBP1, AHCY, AHSG, ALOX15, ALPI, AMFR, ATP2A2, ANGPT1, ANGPT2, APEX1, APOA1, APP, AQP4, AREG, RHOA, ARSA, ARSB, STS, ATD, TNFSF8, CD33, SCARB1, DECR1, CRX, CRYGC, CSTA, CSTB, CTSD, CTSE, CTSL, CYP2A7, DAPK1, DBH, DCN, DDX1, DEFA1, CP, DHCR24, DIO1, DIO2, DIO3, DNASE1, DUSP6, DVL3, EDNRB, EIF4EBP1, ELAVL2, EPHB2, EPHX2, CREBBP, KLF6, CD38, CHRM2, CD47, CD63, CD69, CD74, CD81, CDC6, CDH3, CDK6, CDKN1B, CDKN1C, CDX2, CEACAM5, CHRNA7, COL11A1, CHUK, CIRBP, CKMT1B, TPP1, CCR7, CCR8, LTB4R, CNR2, COL1A1, COL1A2, COL6A1, COL6A2, MEF2D, MEFV, MAP3K5, MFAP4, VWF, WNT1, WNT10B, XDH, YY1, ZFP36, ZNF208, PXDN, SLBP, TFEB, NR4A3, FOSL1, MIA, FZD4, PLA2G10, ULK1, LTBP4, RECK, CUL4A, GPR65, IKBKG, PIR, PLA2G4C, STC2, HSD17B6, AOC3, DDX3Y, VTN, VIPR1, EZR, TP53BP1, TFPI, TGFB3, TGFBR3, THAS, NKX2-1, TM7SF2, CLDN5, TNFRSF1A, TNFRSF1B, TNNI3, TNNT1, TNXB, TPR, VCP, TPT1, NR2C2, TRPC1, TRPC3, TRPC6, TUFM, TULP3, TWIST1, TNFRSF4, TXN, TXNRD1, UROD, ADAM9, RIPK2, TNFRSF10C, IKZF1, KEAP1, MAGI2, MFN2, RBM8A, SRA1, ABCB6, DNM1L, EBI3, WASF2, DHRS2, EIF1, STAM2, TLR6, SOCS5, KLF2, NOD1, YAP1, MCRS1, MERTK, CAP1, CARM1, FBLN5, CIB2, CHERP, ATG7, LILRB1, HDAC4, IKBKE, IL18R1, P2RX6, CES2, NRP1, ASAP2, ARHGEF7, SPHK1, SGPL1, MBD2, WASF1, HSPB3, BTAF1, GPRC5A, SLC16A4, IL1RL1, CELSR1, MTA2, IL32, RPS6KA5, S1PR2, SLIT2, PPIG, CYP7B1, GSTO1, EIF2AK3, AKAP5, CXCL14, AKAP12, TERT, TEAD4, TRBV20OR9-2, PCBP2, NTF3, OAT, OMP, P2RX1, P2RX3, P2RX4, P2RX5, P2RY1, FURIN, PAH, PAK2, REG3A, PCNA, YBX1, PDE4D, PECAM1, PIM1, PITX3, PLA2G2A, PLAT, PLXNA2, PPIA, PPIC, PRKCD, PRKCZ, MAPK7, NT5E, NRL, MAPK13, MTRR, MFGE8, MGMT, MGP, MIF, MIP, KMT2A, MME, MMP8, CD200, MSI1, MSX1, MTHFD1, MUC4, NRAS, MUC6, MYCN, MYD88, MYO1D, MYO1E, NELL1, NFATC3, NFKBIB, NGF, NHS, NOTCH2, NOTCH3, MAPK9, PRNP, TCF4, SPARC, SLC1A5, SLC1A7, SLC2A4, SLC6A8, SLC11A1, SMN1, SMN2, SMPD1, SMS, SOX9, SP3, SPAM1, SPINT1, SGCA, ST2, STAT1, STAT5A, STAT5B, SUV39H1, SVIL, TACR2, ADAM17, TAF1, MAP3K7, TAZ, SERPINA7, SGSH, SFRP5, KLK6, ROS1, PTCH1, PTGDS, PTPN6, PTPRC, PVT1, RARRES2, REN, RIT1, RNF5, ROBO2, ROM1, RORC, RPGR, SFRP2, RPS3, RRM1, S100A4, S100A5, SATB1, CCL3, CCL24, CXCL11, SDC4, SELL, SELP, SET, H3P40
-
Levator Ani Syndrome
Wikipedia
ISBN 978-88-470-1542-5 . ^ a b c Bharucha AE, Trabuco E (September 2008). "Functional and chronic anorectal and pelvic pain disorders" . Gastroenterology Clinics of North America . 37 (3): 685–96, ix. doi : 10.1016/j.gtc.2008.06.002 . ... PMID 15526112 . ^ Rao, SS; Bharucha, AE; Chiarioni, G; Felt-Bersma, R; Knowles, C; Malcolm, A; Wald, A (25 March 2016). ... "Anorectal and Pelvic Pain" . Mayo Clinic Proceedings (review). 91 (10): 1471–1486. doi : 10.1016/j.mayocp.2016.08.011 .
-
Substance Intoxication
Wikipedia
DSM-IV training guide . Psychology Press. pp. 80–. ISBN 978-0-87630-768-7 . Retrieved 27 April 2010 . ^ "Acute intoxication" . World Health Organization . Retrieved 2020-01-31 . ^ Johnson BD, BardhiF, Sifaneck SJ, Dunlap E (2005).
-
Spinal Muscular Atrophy
Wikipedia
Work on developing gene therapy for SMA is also conducted at the Institut de Myologie in Paris [89] and at the University of Oxford . ... Oral salbutamol (albuterol), a popular asthma medicine, showed therapeutic potential in SMA both in vitro [97] and in three small-scale clinical trials involving patients with SMA types 2 and 3, [98] [99] [100] besides offering respiratory benefits. ... American Journal of Physical Medicine & Rehabilitation . 86 (5): 339–45 quiz 346–8, 379. doi : 10.1097/PHM.0b013e31804a8505 . ... Seminars in Pediatric Neurology . 5 (2): 106–15. doi : 10.1016/S1071-9091(98)80026-0 . PMID 9661244 . ^ Tein I, Sloane AE, Donner EJ, Lehotay DC, Millington DS, Kelley RI (January 1995). ... American Journal of Physical Medicine & Rehabilitation . 86 (5): 349–55. doi : 10.1097/PHM.0b013e31804b1d66 .SMN1, SMN2, TRPV4, ASCC1, ATAD3A, DYNC1H1, BICD2, VRK1, UBA1, ASAH1, IGHMBP2, TRIP4, VAPB, EXOSC8, SIGMAR1, TK2, CHCHD10, DNAJB2, SNURF, NAIP, FBXO38, KCNK9, DYSF, PLS3, SNRPN, TBCE, PLEKHG5, LSM2, MAP1B, STMN1, ATP7A, CNTNAP1, GEMIN2, SOD1, IGFALS, ACTB, FUS, TP53, BCL2, SERF1A, ZPR1, IGF1, NCALD, KHDRBS1, SLC9A3R2, GUSBP1, GUSBP3, GTF2H2, MSTN, SCAF11, SYNCRIP, DDX20, ZEB2, SCO2, GUSBP14, DMD, IFI44, H3P33, SERF1B, CARM1, TMEM41B, NEFL, AR, WDR77, HDAC9, TARDBP, COPA, EXOSC3, DCPS, VIM, RPL3, CORO1C, LMNA, PART1, MUC1, HTRA2, LIX1, PTEN, DROSHA, RNPC3, SLCO2A1, CHODL, CARD16, SNCA, SLC25A21, NRXN2, TIA1, CHP1, PLA1A, SLC7A10, AAVS1, ALB, CDK5, DES, CTNNB1, MAPK14, AKT1, GJB1, ETFA, GTF2H1, GARS1, ACVR2B, MORC2, ITGA11, COMP, SETX, KIF1B, NCDN, CREM, CRK, CREB1, IGF2, CD2AP, NUP62, CNTN1, RNF19A, POLDIP2, EIF3K, SGSM3, CHML, KLF15, CD47, GEMIN4, HAPLN1, CSF2, STRAP, HNRNPR, CLOCK, MAD2L1BP, NCOR2, EEF1D, HDAC4, TYMP, SLC23A2, DPP4, ABCB6, RBM7, POP7, GUSBP15, DNM1, SMNDC1, PRMT5, FST, CCN2, AHSA1, CTF1, CELF2, PPARGC1A, CD36, CD40, LEF1, DCTN4, GRAP2, ANXA1, FBXO32, ANG, ALPP, C9orf72, CYP4V2, HCN1, ALPI, GTF2H5, MIR146A, MIR206, MIR223, MIR23A, MIR335, MIR375, MIR431, OCLNP1, HNRNPA1P10, POTEF, AGRP, ADRB2, RNU4ATAC, RNU6ATAC, OCLN, UPK3B, CDCA5, SLC25A46, STARD13, KIDINS220, CAT, CASP8, POP5, VPS54, ARID4B, GEMIN8, WRAP53, CASP3, CANX, CALR, BMP4, CIP2A, APOE, BICD1, ARHGAP22, HCN2, GORASP1, WNK1, BCL2L1, GEMIN6, ATP6V1B2, CCDC8, ARSF, MAGT1, NR1D1, EPHA4, ELAVL4, EMX1, NOS2, OXA1L, PAX3, PAX7, PFN1, PGK1, HTT, PML, PPARG, MAPK1, MAPK10, PSMD4, GTF2H4, PTGS2, PVALB, RAB1A, RELA, RNASE4, BRD2, GTF2H3, RPL9, RPS6KB2, RYR1, S100A1, S100A10, NOS1, NFKB1, HDAC2, MBP, IAPP, IL1B, IL6, IL12A, KIT, LAMC2, HSPG2, LBR, LGALS3, HOXA5, HNRNPA1, MDM2, NEFH, MDM4, MECP2, RAB8A, MAP3K10, MNAT1, NRG1, MYH7, MYOG, HEXA, NCL, NEDD8, S100B, ATXN2, CCL2, KHSRP, BEST1, GAPDH, ZAP70, AIMP2, SPPM, CDK2AP1, COIL, KDSR, FUT1, FOXO1, FCN2, BECN1, UCHL1, SOCS2, CDK5R1, ARHGEF7, SQSTM1, BCL10, ERCC3, ARTN, ERCC2, ERBB4, KLF4, IGF1R, VDAC2, GART, CCL18, STAT5A, CCL21, XCL1, SRSF1, TRA2B, SLC1A3, GSK3B, GRIN2A, GM2A, ACHE, SPP1, SRF, STAT5B, TTN, STATH, SYT1, TBCD, GFAP, PPP1R11, TFRC, GCHFR, GCG, TNF, GATA6, TRH, H3P40
-
Ebola
Wikipedia
In Africa, wild animals including fruit bats are hunted for food and are referred to as bushmeat. [78] [79] In equatorial Africa, human consumption of bushmeat has been linked to animal-to-human transmission of diseases, including Ebola. [80] Although it is not entirely clear how Ebola initially spreads from animals to humans, the spread is believed to involve direct contact with an infected wild animal or fruit bat. [59] Besides bats, other wild animals sometimes infected with EBOV include several species of monkeys such as baboons , great apes ( chimpanzees and gorillas ), and duikers (a species of antelope ). [81] Animals may become infected when they eat fruit partially eaten by bats carrying the virus. [82] Fruit production, animal behavior and other factors may trigger outbreaks among animal populations. [82] Evidence indicates that both domestic dogs and pigs can also be infected with EBOV. [83] Dogs do not appear to develop symptoms when they carry the virus, and pigs appear to be able to transmit the virus to at least some primates. [83] Although some dogs in an area in which a human outbreak occurred had antibodies to EBOV, it is unclear whether they played a role in spreading the disease to people. [83] Reservoir The natural reservoir for Ebola has yet to be confirmed; however, bats are considered to be the most likely candidate. [60] Three types of fruit bats ( Hypsignathus monstrosus , Epomops franqueti and Myonycteris torquata ) were found to possibly carry the virus without getting sick. [84] As of 2013 [update] , whether other animals are involved in its spread is not known. [83] Plants, arthropods , rodents , and birds have also been considered possible viral reservoirs. [1] [29] Bats were known to roost in the cotton factory in which the first cases of the 1976 and 1979 outbreaks were observed, and they have also been implicated in Marburg virus infections in 1975 and 1980. [85] Of 24 plant and 19 vertebrate species experimentally inoculated with EBOV, only bats became infected. [86] The bats displayed no clinical signs of disease, which is considered evidence that these bats are a reservoir species of EBOV. In a 2002–2003 survey of 1,030 animals including 679 bats from Gabon and the Republic of the Congo , immunoglobulin G (IgG) immune defense molecules indicative of Ebola infection were found in three bat species; at various periods of study, between 2.2 and 22.6% of bats were found to contain both RNA sequences and IgG molecules indicating Ebola infection. [87] Antibodies against Zaire and Reston viruses have been found in fruit bats in Bangladesh , suggesting that these bats are also potential hosts of the virus and that the filoviruses are present in Asia. [88] Between 1976 and 1998, in 30,000 mammals, birds, reptiles, amphibians and arthropods sampled from regions of EBOV outbreaks, no Ebola virus was detected apart from some genetic traces found in six rodents (belonging to the species Mus setulosus and Praomys ) and one shrew ( Sylvisorex ollula ) collected from the Central African Republic . [85] [89] However, further research efforts have not confirmed rodents as a reservoir. [90] Traces of EBOV were detected in the carcasses of gorillas and chimpanzees during outbreaks in 2001 and 2003, which later became the source of human infections. ... The widespread bleeding that occurs in affected people causes swelling and shock due to loss of blood volume . [94] The dysfunctional bleeding and clotting commonly seen in EVD has been attributed to increased activation of the extrinsic pathway of the coagulation cascade due to excessive tissue factor production by macrophages and monocytes. [23] After infection, a secreted glycoprotein , small soluble glycoprotein (sGP or GP) is synthesised. ... When a cell is infected with EBOV, receptors located in the cell's cytosol (such as RIG-I and MDA5 ) or outside of the cytosol (such as Toll-like receptor 3 (TLR3) , TLR7 , TLR8 and TLR9 ) recognise infectious molecules associated with the virus. [51] On TLR activation, proteins including interferon regulatory factor 3 and interferon regulatory factor 7 trigger a signalling cascade that leads to the expression of type 1 interferons . [51] The type 1 interferons are then released and bind to the IFNAR1 and IFNAR2 receptors expressed on the surface of a neighbouring cell. [51] Once interferon has bound to its receptors on the neighbouring cell, the signalling proteins STAT1 and STAT2 are activated and move to the cell's nucleus . [51] This triggers the expression of interferon-stimulated genes , which code for proteins with antiviral properties. [51] EBOV's V24 protein blocks the production of these antiviral proteins by preventing the STAT1 signalling protein in the neighbouring cell from entering the nucleus. [51] The VP35 protein directly inhibits the production of interferon-beta. [95] By inhibiting these immune responses, EBOV may quickly spread throughout the body. [49] Diagnosis When EVD is suspected, travel, work history, and exposure to wildlife are important factors with respect to further diagnostic efforts. [ medical citation needed ] Laboratory testing Possible non-specific laboratory indicators of EVD include a low platelet count ; an initially decreased white blood cell count followed by an increased white blood cell count ; elevated levels of the liver enzymes alanine aminotransferase (ALT) and aspartate aminotransferase (AST); and abnormalities in blood clotting often consistent with disseminated intravascular coagulation (DIC) such as a prolonged prothrombin time , partial thromboplastin time , and bleeding time . [96] Filovirions such as EBOV may be identified by their unique filamentous shapes in cell cultures examined with electron microscopy . [97] The specific diagnosis of EVD is confirmed by isolating the virus, detecting its RNA or proteins, or detecting antibodies against the virus in a person's blood. [98] Isolating the virus by cell culture , detecting the viral RNA by polymerase chain reaction (PCR) [6] [23] and detecting proteins by enzyme-linked immunosorbent assay (ELISA) are methods best used in the early stages of the disease and also for detecting the virus in human remains. [6] [98] Detecting antibodies against the virus is most reliable in the later stages of the disease and in those who recover. [98] IgM antibodies are detectable two days after symptom onset and IgG antibodies can be detected six to 18 days after symptom onset. [23] During an outbreak, isolation of the virus with cell culture methods is often not feasible. ... The response to the epidemic then moved to a second phase, as the focus shifted from slowing transmission to ending the epidemic. [190] On 8 April 2015, the WHO reported only 30 confirmed cases, the lowest weekly total since the third week of May 2014. [191] On 29 December 2015, 42 days after the last person tested negative for a second time, Guinea was declared free of Ebola transmission. [192] At that time, a 90-day period of heightened surveillance was announced by that agency.SARS1, ERVK-6, SARS2, ERVK-20, NPC1, ERVW-1, TLR4, TIMELESS, RBBP6, ARHGEF5, HAVCR1, ERVK-32, STAT1, PLAAT4, IFNG, TNF, NEDD4, BST2, GP2, ZNF415, IL10, BAG3, IL6, IFNA13, LSAMP, ATM, ROBO3, ANXA5, IFNB1, IFNA1, PIKFYVE, DDX58, LAMP3, PRKRA, CD209, TPCN2, SLC25A19, IFI30, RUVBL2, LGR6, HAMP, APEX2, MAVS, PELI1, ZC3HAV1, TLR7, CLEC10A, VAC14, MERTK, HSPA14, CASZ1, CPVL, NUSAP1, LPAR3, RETREG1, TLR9, SEPTIN8, MUL1, SMYD3, MIR155, ANO6, GPRC6A, MRGPRX1, CLEC4G, CCL4L1, VHLL, MIR320A, CBLL2, VN1R17P, GPR166P, MIR196B, MIR1246, ECT, LINC01672, OXER1, GPR151, GNPTAB, HAVCR2, IRF9, FUZ, LNPK, HM13, RILP, NTPCR, DISP1, TRIM6, TIMD4, PRDM6, NLRP3, MRGPRX3, MRGPRX4, RLN3, IFITM3, ACP2, ISG15, IL18, HLA-B, HMOX1, HSPA5, HSPD1, IL1B, IL4, IRF1, GP5, IRF7, ITPA, JUN, JUNB, JUND, KPNA1, HLA-A, GALNT1, GTPBP1, EGI, AGRP, ALB, CDSN, CSF2, DAG1, DPAGT1, EIF5A, FOSB, ESR1, FCGR1A, FCGR1B, FOLR1, FOLR2, FOS, KPNA5, LAMP1, LGALS1, ACTB, VTN, XRCC5, USP7, CCDC6, FZD4, RUVBL1, MBTPS1, NHS, RAB11A, ARTN, MYOM2, CD163, SH3BP5, CCL4L2, VEGFA, TYROBP, TSG101, TPI1, TOP1, TMPRSS2, TLR3, THBS1, STAU1, SIGLEC1, SFTPD, SDC1, CCL5, CCL4, PTPA, POMC, PRKN, SOCS1