-
Boomerang Dysplasia
Orphanet
Boomerang dysplasia (BD) is a rare lethal skeletal dysplasia characterized by severe short-limbed dwarfism, dislocated joints, club feet, distinctive facies and diagnostic x-ray findings of underossified and dysplastic long tubular bones, with a boomerang-like bowing. Epidemiology The prevalence of BD is unknown. Clinical description Affected neonates are stillborn or die rapidly after birth and present clinically with severe short-limbed dwarfism, dislocated hip, knee and elbow joints, club feet and proviso born alive have severe cardio respiratory failure. ... Boomerang dysplasia clinically differs from AOI and AOIII because of the boomerang shaped bowing of the femur and occasionally observed encephalocele and omphalocele. Etiology BD results from missense mutations or small in-frame deletions in the FLNB gene reported in exons 2-5, normally expected to translate full length but biochemically abnormal filamin B protein. ... Distinctive radiographic findings are similar to AOI but, BD presents with a more severe deficiency in mineralization, with non-ossification of certain segments of limbs and vertebrates, and a boomerang-like shape of some long tubular bones. ... Antenatal diagnosis The prenatal diagnosis of BD is difficult to ascertain by ultrasound.
-
Postural Orthostatic Tachycardia Syndrome
Wikipedia
In one survey of 138 POTS patients, brain fog was defined as “forgetful” (91%), “difficulty thinking” (89%), and “difficulty focusing” (88%). ... Midodrine [17] [87] [88] [89] Splanchnic–mesenteric vasoconstriction Splanchnic vasoconstriction Octreotide [90] [91] Hypovolemic POTS Synthetic mineralocorticoid Forces the body to retain salt. Increase blood volume Fludrocortisone (Florinef) [92] Vasopressin receptor antagonist Helps retain water, Increase blood volume Desmopressin (DDAVP) [93] Hyperadrenergic POTS beta-blockers (Non-Selective) Decrease sympathetic tone and heart rate. Propranolol (Inderal) [94] [95] [96] beta-blockers (Selective) Metoprolol (Toprol), [87] [97] Bisoprolol [98] [92] Selective sinus node blockade Directly reducing tachycardia. ... Neuroscience and Biobehavioral Reviews . 90 : 174–183. doi : 10.1016/j.neubiorev.2018.04.017 . ... Postural Orthostatic Tachycardia Syndrome (POTS). 215 : 78–82. doi : 10.1016/j.autneu.2018.04.005 .
-
Proximal Renal Tubular Acidosis
Wikipedia
A defect in bicarbonate reabsorption with normal urinary acidification" . Pediatr. Res . 1 (2): 81–98. doi : 10.1203/00006450-196703000-00001 . ... Journal of the American Society of Nephrology . 13 (8): 2160–2170. doi : 10.1097/01.ASN.0000023430.92674.E5 . ISSN 1046-6673 . PMID 12138150 . ^ Gahl WA, Thoene JG, Schneider JA (2002). ... Its resemblance to renal tubular acidosis" . J. Clin. Invest . 47 (6): 1389–98. doi : 10.1172/JCI105830 . PMC 297294 . ... Nephrol . 13 (8): 2160–70. doi : 10.1097/01.ASN.0000023430.92674.E5 . PMID 12138150 . ^ McSherry E (1981).
-
Maple Syrup Urine Disease
Omim
Nomenclature Maple syrup urine disease caused by mutation in the E1-alpha subunit gene is referred to as MSUD type IA; that caused by a mutation in the E1-beta subunit gene as type IB; and that caused by defect in the E2 subunit gene as type II. The phenotype caused by mutation in the E3 subunit is sometimes referred to as MSUD type III (Chuang and Shih, 2001), but that disorder is more commonly referred to as E3 or dihydrolipoamide dehydrogenase deficiency (246900) (summary by Hong et al., 1996). ... There are 5 clinical subtypes of MSUD: the 'classic' neonatal severe form, an 'intermediate' form, an 'intermittent' form, a 'thiamine-responsive' form, and an 'E3-deficient with lactic acidosis' form (246900). All of these subtypes can be caused by mutations in any of the 4 genes mentioned above, except for the E3-deficient form, which is caused only by mutation in the E3 gene (Chuang and Shih, 2001). ... Functional expression studies in E. coli showed that the different mutations had variable expression but no residual enzyme activity. Mutations in the E2 Component Gene In a case of classic MSUD, Herring et al. (1991) identified a 124-bp deletion in the DBT gene encoding the E2 component of the BCKDH complex (248610.0001). ... All patients except 2, 1 with E1-alpha and 1 with an E1-beta mutations, had documented episodes of metabolic decompensation. IQ greater than 90 was observed in 70% of patients. Patients with mutations in the E1-alpha gene tended to have decreased IQs compared to other patients.
-
Head And Neck Cancer
Wikipedia
The role of HPV in the remaining 25-30% is not yet clear. [43] Oral sex is not risk free and results in a significant proportion of HPV-related head and neck cancer. [44] Positive HPV16 status is associated with improved prognosis over HPV-negative OSCC. [45] HPV can induce tumor by several mechanisms: [46] E6 and E7 oncogenic proteins. Disruption of tumor suppressor genes . ... Journal of the National Cancer Institute . 91 (8): 726–8. doi : 10.1093/jnci/91.8.726a . hdl : 2434/520105 . ... The New England Journal of Medicine . 340 (23): 1773–80. CiteSeerX 10.1.1.460.1056 . doi : 10.1056/NEJM199906103402301 . ... Journal of the National Cancer Institute . 97 (7): 481–8. doi : 10.1093/jnci/dji095 . ... Springer Science & Business Media. pp. 257–87. ISBN 978-94-007-5827-8 . Archived from the original on 7 January 2016. ^ Wang YX, Hu D, Yan X (September 2013).EGFR, TP53, PIK3CA, FGFR2, BCL2L1, ERBB3, RARB, XRCC3, RAD51, BAP1, TGFA, GRP, PIK3CB, APOBEC3B, STAT6, VEGFA, FGFR1, FAT1, ERBB2, PRAME, UROD, TYMS, GPX1, MAL, DPYD, CYLD, BCL2, MAPK1, AREG, CEBPA, CSF3, MAPK3, CDC73, NAA25, HELQ, FOXE1, ADH7, MINPP1, ADH1B, HABP2, IL6, PTGS2, AHR, RRM2, XRCC1, IL13, PTGS1, CCND1, CA9, CYP1A1, STAT3, S100A2, TCF21, ICAM5, STAT5B, SDHC, TNF, STAT5A, NAT2, TP53BP1, VIM, HOTAIRM1, MTRNR2L12, MIR210, MIR204, MIR148A, ABCC10, MRPL11, ING3, CYCS, TLR7, ING4, TMEM97, BBC3, HPSE, ROBO1, DLGAP1, IFITM1, GJB6, MSH2, RAD17, MAPK6, GJB2, FGFR4, EPOR, EPO, EPHB3, EPAS1, EIF4E, EGF, DSG3, DDIT3, CYP1B1, CD44, CD4, BRAF, BRCA1, BMI1, B2M, AR, ALDH1A1, GNAS, GSTM1, GSTP1, MKI67, PCYT1A, FURIN, NOS2, NME1, MTRR, MTR, ABCC1, MLH1, MET, HIF1A, KDR, IL13RA2, IL2RA, IGFBP3, IGF1R, IGF1, HLCS, HLA-C, CERNA3
-
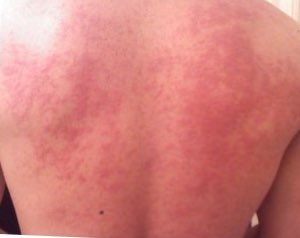
Food Allergy
Wikipedia
Food allergy affects as many as 5% of infants less than three years of age [72] and 3% to 4% of adults. [73] [80] The prevalence of food allergies is rising. [70] [81] [82] Food allergies cause roughly 30,000 emergency room visits and 150 deaths per year. [83] Society and culture [ edit ] Whether rates of food allergy are increasing or not, food allergy awareness has definitely increased, with impacts on the quality of life for children, their parents and their caregivers. [84] [85] [86] [87] In the United States, the Food Allergen Labeling and Consumer Protection Act of 2004 causes people to be reminded of allergy problems every time they handle a food package, and restaurants have added allergen warnings to menus. ... Despite all these precautions, people with serious allergies are aware that accidental exposure can easily occur at other peoples' houses, at school or in restaurants. [89] Food fear has a significant impact on quality of life. [86] [87] For children with allergies, their quality of life is also affected by actions of their peers. ... Food allergens prioritized in labeling laws by country Food US Canada UK Australia EU peanuts Yes [92] Yes [93] Yes [94] Yes [95] Yes [96] tree nuts Yes [92] Yes [93] Yes [94] Yes [95] Yes [96] milk Yes [92] Yes [93] Yes [94] Yes [95] Yes [96] eggs Yes [92] Yes [93] Yes [94] Yes [95] Yes [96] fish Yes [92] Yes [93] Yes [94] Yes [95] Yes [96] shellfish Crustaceans only [92] Crustaceans and molluscs [93] Crustaceans and molluscs [94] Yes [95] Crustaceans and molluscs [96] soy Yes [92] Yes [93] Yes [94] Yes [95] Yes [96] wheat Yes [92] Includes triticale [93] Included under gluten [94] Yes [95] Included under gluten [96] sesame seeds Voluntary [97] Yes [93] Yes [94] Yes [95] Yes [96] mustard No Yes [93] Yes [94] No Yes [96] sulphites (not a true allergy ) No Yes [93] Yes [94] No Yes, >10 mg/kg [96] gluten (not a true allergy) No Yes [93] Yes [94] No Yes [96] celery No No Yes [94] No Yes [96] lupin No No Yes [94] Yes [95] Yes [96] There are no labeling laws mandating declaration of the presence of trace amounts in the final product as a consequence of cross-contamination, except in Brazil. [12] [98] [99] [100] [101] [102] [10] [11] Ingredients intentionally added [ edit ] In the United States, the Food Allergen Labeling and Consumer Protection Act of 2004 (FALCPA) requires companies to disclose on the label whether a packaged food product contains any of these eight major food allergens, added intentionally: cow's milk, peanuts, eggs, shellfish, fish, tree nuts, soy and wheat. [98] The eight-ingredient is list originated in 1999 from the World Health Organisation Codex Alimentarius Commission. [10] To meet FALCPA labeling requirements, if an ingredient is derived from one of the required-label allergens, then it must either have its "food sourced name" in parentheses, for example "Casein (milk)," or as an alternative, there must be a statement separate but adjacent to the ingredients list: "Contains milk" (and any other of the allergens with mandatory labeling). [98] [100] The European Union requires listing for those eight major allergens plus molluscs, celery, mustard, lupin, sesame and sulfites. [99] In 2018, the US FDA issued a request for information for the consideration of labeling for sesame to help protect people who have sesame allergies. [103] A decision was reached in November 2020 that food manufacturers voluntarily declare that when powdered sesame seeds are used as a previously unspecified spice or flavor, the label be changed to “spice (sesame)” or “flavor (sesame).” [97] FALCPA applies to packaged foods regulated by the FDA, which does not include poultry, most meats, certain egg products, and most alcoholic beverages. [11] However, some meat, poultry, and egg processed products may contain allergenic ingredients. ... Gastroenterology . 148 (6): 1120–1131.e4. doi : 10.1053/j.gastro.2015.02.006 . ... J Investig Allergol Clin Immunol . 15 (2): 86–90. PMID 16047707 . ^ Sicherer 2006 , p. 189 ^ Turnbull JL, Adams HN, Gorard DA (2015).
-
Side Effects Of Bicalutamide
Wikipedia
4 months Interstitial pneumonitis Death Kawahara et al. (2009) 8 78 years Male 80 mg/day 8 months Interstitial pneumonitis Recovered Masago et al. (2011) 9 77 years Male ? ... The median age of the patients was 73.5 years (range 59 to 91 years), and median duration of bicalutamide exposure was 7.5 weeks (range 1 to 312 weeks). ... "The role of antiandrogen monotherapy in the treatment of prostate cancer". BJU Int . 91 (5): 455–61. doi : 10.1046/j.1464-410X.2003.04026.x . ... "The role of antiandrogen monotherapy in the treatment of prostate cancer" . BJU International . 91 (5): 455–61. doi : 10.1046/j.1464-410X.2003.04026.x . ... Urology . 47 (1A Suppl): 70–9, discussion 80–4. doi : 10.1016/s0090-4295(96)80012-4 .
-
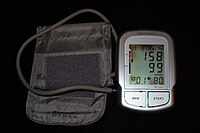
Hypertension
Wikipedia
A chest X-ray or an echocardiogram may also be performed to look for signs of heart enlargement or damage to the heart. [23] Classification in adults [ edit ] Classification in adults (Persons with systolic and diastolic in different categories are assigned to the higher category. [7] ) Category Systolic , mmHg Diastolic , mmHg Hypotension < 90 < 60 Normal 90–119 [7] 90–129 [83] 60–79 [7] 60–84 [83] Prehypertension (high normal, elevated [7] ) 120–129 [7] 130–139 [83] [84] 60–79 [7] 85–89 [83] [84] Stage 1 hypertension 130-139 [7] 140–159 [83] 80-89 [7] 90–99 [83] Stage 2 hypertension >140 [7] 160–179 [83] >90 [7] 100–109 [83] Hypertensive crises ≥ 180 [7] ≥ 120 [7] Isolated systolic hypertension ≥ 140 [7] < 90 [7] Isolated diastolic hypertension [85] [86] < 140 ≥ 90 In people aged 18 years or older hypertension is defined as either a systolic or a diastolic blood pressure measurement consistently higher than an accepted normal value (this is above 129 or 139 mmHg systolic, 89 mmHg diastolic depending on the guideline). [5] [7] Other thresholds are used (135 mmHg systolic or 85 mmHg diastolic) if measurements are derived from 24-hour ambulatory or home monitoring. [73] Recent international hypertension guidelines have also created categories below the hypertensive range to indicate a continuum of risk with higher blood pressures in the normal range. The Seventh Report of the Joint National Committee on Prevention, Detection, Evaluation and Treatment of High Blood Pressure (JNC7) published in 2003 [27] uses the term prehypertension for blood pressure in the range 120–139 mmHg systolic or 80–89 mmHg diastolic, while European Society of Hypertension Guidelines (2007) [87] and British Hypertension Society (BHS) IV (2004) [88] use optimal, normal and high normal categories to subdivide pressures below 140 mmHg systolic and 90 mmHg diastolic. ... Hypertension is classified as "resistant" if medications do not reduce blood pressure to normal levels. [27] In November 2017, the American Heart Association and American College of Cardiology published a joint guideline which updates the recommendations of the JNC7 report. [89] The 2020 International Society of Hypertension guidelines define hypertension based on office blood pressure ≥140/90 mmHg or home monitoring blood pressure ≥135/85 mmHg, or 24-hour ambulatory blood pressure average ≥130/80 mmHg (daytime average ≥135/85 mmHg or nighttime average BP ≥120/70 mmHg). [90] Children [ edit ] Hypertension occurs in around 0.2 to 3% of newborns; however, blood pressure is not measured routinely in healthy newborns. [33] Hypertension is more common in high risk newborns. ... High blood pressure must be confirmed on repeated visits however before characterizing a child as having hypertension. [91] Prehypertension in children has been defined as average systolic or diastolic blood pressure that is greater than or equal to the 90th percentile, but less than the 95th percentile. [91] In adolescents, it has been proposed that hypertension and pre-hypertension are diagnosed and classified using the same criteria as in adults. [91] The value of routine screening for hypertension in children over the age of 3 years is debated. [92] [93] In 2004 the National High Blood Pressure Education Program recommended that children aged 3 years and older have blood pressure measurement at least once at every health care visit [91] and the National Heart, Lung, and Blood Institute and American Academy of Pediatrics made a similar recommendation. [94] However, the American Academy of Family Physicians [95] supports the view of the U.S. ... Annals of Internal Medicine . 162 (3): 184–91. doi : 10.7326/M14-0773 . PMID 25531552 .PPARG, APOA1, HSD11B2, MMP2, NOS3, ATP2B1, APOE, NR3C1, ECE1, GJA1, HIF1A, IL1B, CAV1, CAT, CYBA, AVP, NOX4, NOS2, CCL2, AGT, NPPB, ADRB1, REN, RGS2, AGTR2, KLK1, NPPA, NPR1, TNF, ADD1, CD36, MMP9, CRP, ACE, ACE2, TGFB1, LEP, IGF1, CYBB, EDNRA, PTGS2, EDN1, HMOX1, SERPINE1, AGTR1, UCP2, VWF, TH, UTS2, INS, HP, BDKRB2, PPARA, SOD1, GLP1R, ATP1A1, RGS5, AR, KCNMB1, SOD3, SOD2, SLC12A2, TRH, AQP1, GRK2, GJA5, TIMP1, OLR1, MTOR, VCAM1, BCL2, GPX1, ADRA2A, RELA, JUN, GSK3B, FOS, GAL, PRKCD, ADCY5, IKBKB, WNK1, WNK4, OXT, ATP1A2, ELN, HRH3, SCNN1B, SCNN1G, UMOD, ANXA1, SLC9A3R2, SCNN1A, TP53, GNAS, GUCY1A1, GCH1, CBS, FN1, COL3A1, PTH, POMC, COL1A1, GSTT1, TLR4, EPO, GNB3, CALCA, LPL, NEDD4L, SLC12A3, F12, VDR, CYP1A1, CYP11B1, ADIPOQ, KNG1, IL6, ALB, LEPR, STK39, GCG, ENG, HSD11B1, PLAT, ICAM1, FBN1, LOX, SELP, DRD2, SLC6A2, CCN2, AHR, EGFR, PLG, THBD, NCF1, C3, PRKG1, OXSR1, SH2B3, CXCR2, INPPL1, TRPC3, ERAP1, CMA1, ORAI1, PTPN1, CACNA1C, CLCNKA, GJA4, TGM2, CETP, AOC3, ALOX15, SLC6A18, STIM1, GCLC, TACR1, GSTP1, CRHR2, HTR2B, ALAD, VAV3, PTEN, DUSP5, KCNJ1, F11, IER3, MYO6, TRPC5, STK11, AOC1, UCN, ATP2B3, PROC, RPS6KB1, ITM2B, CELA2A, CHGA, CACNA2D1, NPTN, RALBP1, PPBP, GSTM1, NPY, ADRB2, ADRA2B, ADRA1A, GJC1, GSTT2, ADM, EDN3, CXCL2, ESR2, ATP6AP2, SLC9A3, MME, MYH7, MDH1, GNRH1, TPM1, APLN, FXYD2, LGALS3, ABCC1, CYP4A11, MYD88, GSTA5, ATOX1, REG3G, KL, LHB, MC2R, SLC5A2, NFE2L2, LTF, IL10, GP1BA, KDR, AGER, XDH, F3, MAS1, EDNRB, ARG1, CYP19A1, PNMT, NOS1, SLC8A1, CLOCK, AKT1, GSTM2, ACSM3, COMT, CHI3L1, IL1RN, DBH, MFN2, F11R, UTS2R, KCNQ1, APP, ADD2, AVPR1A, EPHX2, KCNA5, MMP7, CCR5, FADS1, HAVCR1, CDKN2A, SST, HDAC6, RAG1, SLC2A1, ALOX5, ADA, AQP2, FGF2, BDKRB1, ARG2, ARID1B, HSPD1, BECN1, EGLN1, NQO1, CRHR1, CNR1, ADAMTS16, BAX, MIF, CASP3, PDGFRB, AKT2, GNAI2, MAPK8, HTR1B, PRKCA, PTK2B, SRF, KYNU, CPS1, GRK5, AGTRAP, SLC6A3, CCL28, KCNJ11, TAC3, YWHAZ, HDAC8, SLC4A4, HSD3B1, TRHR, STS, UCP3, LRP2, CYP11A1, RHOA, NCF2, CTSC, ALOX12, NPY1R, SLC6A19, PLCD1, PRKCB, POSTN, CYP4F12, PDGFRA, IGF1R, RAMP2, F2R, SLC9A3R1, S1PR1, ABCC9, OPA1, IGFBP2, NISCH, NGF, LIPG, F7, KCNJ8, CLDN16, ENO1, ITGAV, KCNAB1, ATG7, CYP11B2, CYP17A1, ABAT, CORIN, AQP4, MTPN, HSPG2, CUL3, APOB, DUSP1, RFFL, DRD3, ACTC1, DYNLL1, TJP2, HSD17B4, CYP4A22, CD82, SQSTM1, FADD, GRK3, ADD3, GABBR1, ADORA1, RESP18, BAD, GNAI3, TNFRSF11B, ARHGDIA, H2AX, SLC2A3, NFKBIA, NGFR, KLHL3, RNPEP, RGS4, MAP1LC3A, CASP8, CASP9, RAD51, ATF2, OXTR, QDPR, PTPRJ, CCNE1, PTK2, PDGFB, ENTPD2, SERPINE2, PIK3R1, PKD1, PRKCE, MYOD1, COX5B, NR3C2, TACR3, FAH, ARRB2, HSD3B2, EP300, MC4R, ATP2A3, ATP5F1A, AVPR2, CDO1, NR2F2, TBXAS1, EIF4G2, MSX2, NUDT1, MYH9, SRC, COX2, BMPR2, BRCA1, KCNK3, LDLR, KCNJ5, ENPEP, PKD2, NOTCH3, FLT1, ACVRL1, PDE3A, FGF5, HFE, CACNB2, CFH, COX1, TGFBR2, JAK2, TRPC6, CDKN2B-AS1, ABCB6, ABCC6, CASZ1, STOX1, ENPP1, CACNA1H, PCSK9, ARHGAP42, LMNA, SMAD3, NT5C2, CFHR3, CCND1, SERPINA6, LYZ, CTLA4, ARMC5, PKHD1, FMR1, TET2, NOTCH1, NF1, MYH11, ANTXR2, B2M, FOXF1, MGP, TGFBR1, ALMS1, VHL, WT1, FMO3, CNNM2, ULK4, MKKS, GLA, PGPEP1, TGFBR3, RBM47, PLCE1, NFU1, TNFRSF11A, NOD2, TGFB3, TBX2, KRT8, SMAD6, CD46, MEF2A, HMBS, MOV10, L3MBTL4, ND1, MTTP, CFHR1, MYLK, ANGPTL6, FURIN, PLIN1, PMM2, PRKACA, PRTN3, ABCG8, IFIH1, GATA5, ATXN2, GRK6, RPTOR, H19, CD2AP, FGA, CYP21A2, CACNA1D, ACAT1, NKX2-5, CLCN2, ACTN4, COL4A1, CALR, BRCA2, CPOX, SRGAP3, OFD1, SESN2, MIR21, LRIG2, OR5B12, PDE11A, FGD4, BORCS7, TTC8, ADRB3, SLC25A11, DNAAF1, GPR101, ZNF831, ITGA8, PLEKHG1, ELP1, DBP, AGAP1, COL4A3, ZNF536, PAM16, BNC2, PDE8B, CRIP3, RAPGEF5, LRRC10B, PSTK, DECR1, APOL1, EML6, VANGL1, BMS1, CYP4F2, ARL6, BBS5, DIS3L2, PDE5A, MACROD2, STN1, EDA, PIK3CG, SMARCAL1, ERCC8, TRIM28, ND5, COQ7, SUGCT, MICALL2, XRCC4, ZMPSTE24, CFI, IDUA, DARS2, BBS10, ABCB1, PGF, LINC02875, SLC2A10, TMEM212, MKS1, INSR, MAFB, DPEP1, PLEKHH2, FAM131A, MFAP5, CEP290, IL17A, FGF23, TMEM70, BAG6, DPP4, FUZ, SGCZ, KRT18, WDPCP, COL4A4, COL4A5, TRNQ, TRNS1, TRNS2, SLC33A1, DNAJB11, TRNV, TRNW, MUC1, TAF1C, SELENBP1, P4HA2, TRIP13, SLC52A3, ALDH2, PHF21A, PGR-AS1, MYO9B, DYRK1B, ADA2, BBIP1, TMEM67, ND6, MTHFR, CYTB, COX3, TRNE, AIP, TRNC, TRNF, MMP14, TRNH, CST3, USP8, CERS5, TRNK, MPL, TRNL1, CEP19, BORCS7-ASMT, NLRP3, COL5A1, LPA, LZTFL1, LMX1B, CYP2J2, COL5A2, CYP3A4, NOTCH2, HOTTIP, ANGPT2, CYP3A5, MLXIPL, KLK4, NPHP1, TUBB6, IQCB1, PRRX1, BANF1, LRP6, NDUFS3, SMAD4, MAT2A, TARID, APLNR, ZDHHC2, KLHL29, NFATC2, NFIX, CYP2B6, PTPMT1, ADAMTSL4, CYP2C19, LYN, ANGPT1, APBB1IP, CCHCR1, LSP1, WRN, YES1, WNT2B, ERCC4, MTCO2P12, BBS2, GRK4, GPR39, CDH23, BBS4, GPR20, PTPN22, SWAP70, KIF1B, IFT172, ALX4, CAD, HPSE2, NSMCE2, HOXA-AS2, TRAF3IP1, GPT, TRIM32, SLC39A8, POU6F2, REST, ACACA, CAPZA1, ERCC6, BSCL2, TGFB2, ESR1, ABCG5, EIF4BP8, BBS1, CEP164, PON1, RNU1-4, EXT2, F2, FGF21, GML, LDLRAP1, EDA2R, LRRC7, G6PC, ZNF318, FRMD3, SLC12A1, BST1, KCTD1, ZFAT, COPD, SLC37A4, CC2D2A, NPHP4, WDR35, LEMD3, FOXE3, ARHGAP31, HECTD4, LINC00862, RGL3, GPC3, GH1, TRPV4, FES, VPS51, CFB, CUX2, RPGRIP1L, STK3, SGK1, FGFR2, LARS2, SHBG, RETN, SIRT1, SLC12A9, GBA, WDR19, NPHP3, HSPB8, ND4, HOXA3, BBS9, HLA-DPA1, HLA-DPB1, SLC22A7, CCDC28B, SDCCAG8, TM7SF2, HMGCS2, CDH18, PRKAR1A, ACTA2, PTPN11, C8orf37, TNXB, BBS12, TMEM237, PTGIS, HOXB7, HOXA11, FGD5, ATM, YY1AP1, TLR1, HBB, THPO, OSGEP, BBS7, IKZF5, INF2, POR, IFT27, RARRES2, INVS, POU3F4, AFG3L2, HGD, SLC52A2, VEGFA, ACTB, PPARGC1A, CLDN19, SPP1, FTO, CSK, CTNNB1, NOX1, HCRT, LCN2, HGF, GDF15, GSTK1, RNLS, DDAH1, CYP2C9, GABPA, PTX3, CSF2, F5, SLCO6A1, PIK3CD, GPR42, CABIN1, MAT2B, HSPA4, LPAR2, MMP3, PIK3CB, ACKR3, ANO1, CASR, PIK3CA, IL1A, RAC1, BEST1, HMGB1, NPR3, BDNF, BRS3, CLCNKB, THBS1, CXCR6, TXNRD3, MAPK3, LAMC2, EPAS1, SGSM3, SIRT3, PPIG, FAAH, IFNG, SSTR4, ARTN, TLR2, TRPV1, CD34, MOK, PITX2, ROS1, APRT, APOC3, BMP4, STAT3, SLC26A4, KLK3, PTGS1, ADAM17, PDC, SPHK1, MIR155, OR10A4, IRS1, SETD2, ROCK2, GPER1, MFAP1, AGRP, HPGDS, OSR1, IL18, DRD1, COL18A1, TNFRSF1B, ACR, MAPK1, CD59, CHDH, STK24, CYP2D6, ANPEP, ARSA, PPIA, MIR214, MIR29B1, APOA5, CHST3, MIR29A, P2RX7, DYM, ADORA2B, MIR204, IL4, CXCL8, ADRA1D, EGF, MIR145, MLC1, CRH, EMILIN1, RBP4, MTRR, SELPLG, TAC1, CYP1B1, CYP1A2, HDLBP, SELE, DUOX2, FAS, LOC102724197, PER1, ADM2, CX3CL1, MIR223, SLBP, SLC6A4, GCK, BTBD8, IGF2, SLC7A1, G6PD, GPR151, PPARGC1B, PLA2G7, MRGPRX1, LPAR3, SERPINA5, DAPK2, CCR2, KLF15, STAM2, MIR29B2, HLA-B, FLNA, CAST, ATP5PF, OXER1, OGA, AZU1, HEBP1, EIF2AK4, VN1R17P, S100A6, PRKAA2, RAMP1, S100A12, TRPM8, CRY2, TOR2A, PHACTR1, GPR166P, SCN10A, HTR2A, LGR6, GPRC6A, TLR3, PRKD1, ND2, FZD4, MRGPRX4, PIEZO1, MINDY4, KCNMA1, DUOX1, DAPK3, APEX1, CTH, CRY1, MRGPRX3, GAL3ST1, PTGES, AFP, CYP2C18, MYOCD, DLD, GGTLC1, PAH, HAP1, MIR146A, MBD2, CD44, UCP1, TXN, CD40LG, SULT1E1, ABCA1, TSC1, CX3CR1, ISYNA1, C5AR1, SERPINA3, PSMD9, MMP1, MAPK9, CLIC4, GGT1, NR1I2, S100A4, ROCK1, SUCNR1, SHMT1, SLC4A5, GDF2, CDH13, RYR2, PRKAB1, LIPC, PRKAA1, FGF1, ARHGEF40, CRK, DDR1, MIR130A, CAPN10, FABP3, FASN, BTN2A1, MBL2, BCHE, ACCS, CST12P, COX8A, FABP2, GRAP2, ASIC1, POLDIP2, BMP10, JAG1, CYP2C8, MIR17, IL33, CRMP1, VNN1, TNNI3, SLC9C1, UTRN, CD14, INSRR, FGB, TBX4, CXCL12, ATP1B1, ACSS2, AHSA1, CENPJ, PRCP, DDAH2, PADI4, MAPK14, EHMT1, BSND, PHA2A, SLPI, DENR, AIMP2, HNP1, EDN2, SLC9A1, PLA2G15, DNM1L, MALAT1, CLU, ADCYAP1, SESTD1, DMD, PHEX, SLC25A3, PFN1, PDE4D, JPH3, CNN1, PARP1, PECAM1, DIO2, TESC, P2RX3, MIR143, NUCB2, MPO, ITGB2, IAPP, SLC2A5, SLC35A1, DNMT1, SIK1, ADIPOR1, MPI, RNF19A, HTN3, VEGFC, VIP, EGR1, LBP, LAD1, CIMT, TNC, YAP1, PART1, SERPINF2, FOXO1, SLC5A4, RNF213, ADRA2C, CMKLR1, COMMD5, DHX40, ID2, HDAC9, MIR424, SMUG1, SULF2, USE1, HNF1A, SNAI1, AKR1B1, ZC3HC1, CLTRN, BCL2A1, SRM, ST2, BGLAP, ABCC8, POU2F3, TXN2, NOG, ULK3, SIRT6, NNT, PRDX2, DST, MIR518C, TBX1, SERPINA7, BMP2, ATP2B2, SV2C, IGF2BP2, TMED7-TICAM2, GGT2, GGTLC3, CPQ, WNT5A, GNA14, APC, VCL, SORBS1, UGT1A1, UQCRFS1, SLCO1B1, PDLIM5, GOSR2, ANXA5, SCGB1A1, ARSD, CD274, CNBP, PXDN, RHOBTB1, HBHR, FOXP3, ADRA1B, NR4A3, IL22, NOX3, ZGLP1, TMED7, GGTLC4P, TRPM7, ADORA2A, ADH1B, DEGS1, TUG1, CXCR4, NAMPT, SLC4A7, TWIST1, TFAM, TTR, MCM3AP, ALDH7A1, THBS4, B3GAT1, RN7SL263P, ATR, AKAP10, AMH, ABO, GPR182, SLC17A5, TGFA, MMRN1, SEPHS1, ERAL1, DKK1, FBXO8, ADAMTS13, SDS, C1QL1, TPT1, IMPACT, TSHR, TRPM2, GGTLC5P, SERPINC1, EBP, CRISP2, ATP2A2, MIR510, FOXRED1, NFAT5, NES, ANGPTL3, TNFRSF1A, TMSB4X, ABCG2, ARNTL, NECTIN1, PLN, MIR142, PRSS8, CD40, NLRC3, IL16, PRL, PRKCH, HBG2, HCRTR1, CD68, CD69, SLC9B2, ELK3, MS, PPP3R1, ARID3A, PTPA, TICAM2, ENTPD1, PSEN1, FSD1, ZBTB7C, AGXT2, CD5L, CD28, IFNB1, ETS1, PTGER4, PTGER1, PTGDS, COL17A1, FBL, ELANE, SETD3, PRSS55, BPIFB4, UBE2Z, CD38, FNDC1, NFYA, SMARCA4, ETV3, MIR23A, SERPINB2, PLEKHA7, SPZ1, MIR150, MIR221, CPB2, PEAR1, MIR22, TLX2, CCNL2, PDK1, CLCN6, NEFH, PDE3B, HPT, RETNLB, PCSK1, EPHB1, MIR25, NOX5, MIR26B, CDK6, SLC29A1, CDKN1A, NRG1, PPARD, EPHB6, FSD1L, EPHB2, HT, P2RY6, HLA-DRB1, PCSK6, MIR27B, PLXNA2, LRPAP1, MIR27A, PAEP, NFIL3, GSTM3, CCL5, SCT, SLC2A2, LGALS2, SHC1, HJV, IDE, SFTPC, SFRP5, GATA4, GCGR, GDNF, SDHB, SCPEP1, CCL21, GEM, GFAP, NPPC, CTSB, MAP6, SLC2A4, NPNT, HCCAT5, SLC22A2, SEMA6A, LRP1, BSG, BTF3P11, CYP4F3, SMAD2, FOSL2, MAOB, LIPA, FSHMD1A, TRIB3, FSHR, ARHGEF25, C1QTNF1, SLC3A1, GHRH, NPL, GHSR, FBRS, CAPN1, KCNJ13, GPR35, DHFR, RFC1, CYB5R3, RENBP, NTS, EFNB3, SLC39A12, OPN1LW, RCN2, CSMD1, GRN, CXCL1, GSR, MIR34A, OGG1, MIR192, DCN, NT5E, MIR126, LCT, IGAN1, SAI1, CTSL, KLKB1, RYR3, RYR1, MDK, LINC01194, GOLPH3, EFEMP1, GNA12, FABP1, CBLN2, SIK1B, SENP1, PADI1, CD300LG, APCDD1, TRPM5, GRHL1, GNPDA2, PSAT1, TDRD9, SLC26A9, MIR107, SLC22A12, MIR103A2, MIR100, MRPL4, ATL1, MIRLET7B, FIS1, GSTO2, CYP2R1, SOST, MAP3K15, CYCSP25, FOPNL, NDUFS7, LYPLAL1, BOLA3, TAS2R13, IL31RA, CD164L2, TRIM69, TRPM6, ARMS2, SRXN1, CACUL1, ENHO, KLK12, KCNIP3, RHOD, MYLIP, XPO5, TICAM1, CNTNAP2, EPHA6, SDK1, ADGRE4P, MARCHF8, LOC107987479, ZNF260, RXFP4, ATOH7, DLEU7, HYLS1, TSPAN33, NUPR1, ITGB1BP2, CPP, FOLH1B, SLC36A1, PABPC1, DNM3, GLCE, DROSHA, SLC29A4, NPW, LDOC1, PGP, IFNL2, IS1, BRD1, COL6A5, SLC24A2, KLK9, QPCT, GCA, CASC2, EBF3, FNDC5, VAMAS6, ADGRA2, SH2B1, CMYA5, FLCN, DCP1B, RLS1, LCE1C, RABGEF1, PCSK1N, AMOT, CBSL, SLC2A12, MCAT, CNTN4, SLCO4C1, RMDN2, FAM78B, REM1, IL23R, MOB3C, FLVCR1, RGCC, SLC35F3, PACERR, PAQR7, XIRP1, GPR162, CRPP1, NLRP6, GOLGA6A, MCIDAS, SNX5, HCAR1, CAVIN4, ADAMTS18, DDX53, CERNA3, TCEANC, LINC02210-CRHR1, PDCD4, SLC30A8, LCE1A, BMPER, RMDN1, CNTN5, C1QTNF6, PLXNA3, MTMR9, LGR4, WNK3, SAGE1, MIR30A, ACD, SLC30A10, POU5F1P4, SLC26A6, MIR31, MIR96, RNPC3, MIR33A, P2RY12, SYBU, POU5F1P3, MFSD1, PDIA2, MIR34B, GORASP1, STYK1, MIR585, MIR593, VKORC1, TTC21B, GSTCD, MSTO1, MCPH1, MIR296, PNPO, RMDN3, MIR637, ADIPOR2, SAP130, RCBTB1, MIR621, BHLHE41, PPP6R3, SLC29A3, MIR608, DRAM1, CARD14, TMEM38A, SLC48A1, MIR509-1, OXR1, HAMP, ZNF77, SLC2A9, MIR202, NGB, GRHL3, PLF, CISD2, SPHK2, MEPE, FAM20C, MIR506, MIR410, MIR425, MIR429, IBD7, PCBP4, MIR451A, BEND3, MIR431, VARS2, KCNQ5, IL21, MIR495, SLC5A7, MIR503, ARHGEF28, MIR499A, MIR98, MIR17HG, XYLT2, XYLT1, VPS11, TRIM67, PAG1, SLC28A3, SRR, GAS5, ANGPTL8, MIR328, PDGFC, BTNL2, SULT1A4, TRPV5, TREML2, MAP9, ANGPTL4, CCNQ, PPP1R14A, IL23A, MUC16, MIR132, MIR140, GHRL, PPR1, ASIC5, ORAI3, SLCO1C1, TSLP, CA14, TRIM47, GPR88, SOX18, MCU, WNT4, KNSTRN, TERF2IP, UCN2, PRRT2, OSER1, MRAP2, SFMBT1, HSPA14, MIR10A, C1QTNF3, PIPOX, GOLM1, MIR122, KCNK9, LOC102723407, CYGB, TLR8, TMEM54, MEX3C, BIRC6-AS1, GLCCI1, TAOK3, MIR4513, ANGPT4, IL17F, PRKAG2, SSH1, TAF7L, MIR222, CDKAL1, MIR200A, MIR200B, WBP1L, HDHD2, MIR206, EMILIN2, MIR1224, MIR543, MIR210, COL21A1, PTPN5, TUBB1, COASY, PDGFD, MIR212, FHOD3, RUBCNL, PTGES2, SVEP1, SLC52A1, CHCHD5, ERCC6L, HOPX, MIR19B1, SLX1A-SULT1A3, PPP1R9B, LINC01672, MIR185, MIR190A, ROBO4, SPATA6, KIF2B, MIR195, NDFIP2, MIR199A1, MIR199A2, CROT, ROPN1, ZGPAT, ALL2, NT5C1A, ALL1, TRPM4, A2M, SELL, AMACR, HNRNPD, HSPA1B, HSPA1A, AGFG1, HPX, HPV18I2, HPCAL1, HNRNPF, SLC29A2, HSPA2, HNF4A, NR4A1, HMOX2, HMGCR, HLA-C, HLA-A, UBE2K, HSPA1L, HSP90AA1, HDAC1, IGFALS, IL4R, IL2RA, IL2, IL1R1, CCN1, IGFBP3, IGFBP1, IFNA13, HTN1, IFNA2, IFNA1, IFI27, IDH2, IDH1, ID1, HTR1E, HDAC2, HBG1, CXCR1, FPR2, GATA3, GALNT2, GAD2, XRCC6, GAST, NR5A1, FXN, FPR1, GHR, FOSB, FOLH1, FLT4, FLI1, FOXO3, FOXM1, FOXC1, GC, GIP, HBA2, GSTM5, HBA1, GYS1, GYPE, GYPB, GYPA, GUCY1B1, GUCA2B, GSTA1, GLI2, GRIA1, GPX3, CXCR3, NPBWR1, GPI, GOT2, GLO1, IL5RA, IL10RA, OGN, MMP8, MTHFD1, MT1A, MSR1, MPV17, MPST, MMP12, MMP10, KMT2A, TRNI, MAP3K10, CXCL9, MET, MEST, MAP3K5, MEFV, ADAM11, MTR, MVD, MBP, NFATC3, NPY2R, NPHS1, NPC1, NPAS2, NOVA2, NHS, NFKB1, NEDD9, MX1, NDUFS4, NDUFA10, PPP1R12A, MYOC, MYL3, MYL2, MYH6, MC3R, MAT1A, IL10RB, PDX1, ITGB3, ITGAX, ITGAM, ITGA2, ISG20, IRF1, IRAK1, INHA, ITPR1, IMPA1, ILK, ILF3, IL15, IL13, IL12B, IL11, ITGB8, ITPR3, MAOA, LCN1, SMAD9, SMAD7, LSS, LRP5, LLGL2, LHCGR, LGALS1, KRT16, JUNB, KRT5, KIT, KIR3DL1, KIR2DS5, KCNN3, KCNH1, JUND, FHL2, FHL1, FGFR4, BMP7, C4BPA, C3AR1, BUB1B, KLF5, BRAF, BNIP1, BMPR1A, BGN, C5, BCL6, BCL2L1, BAAT, ATP7A, ATP6V0A1, ATP5F1B, ATP4A, CAPN5, CA2, ATP12A, CAV3, CDSN, CDR1, CDKN2B, CDH2, SCARB2, CD86, RUNX1T1, CASQ2, CACNA1A, CASP1, CAMK2A, CAMK4, CALD1, CALCR, CALB2, SLC25A20, ATP2B4, ATIC, FGFR1, ACVR2A, AMBP, ALOX15B, ALOX5AP, AHSG, NR0B1, AGXT, ADCYAP1R1, ACP3, AMY1B, ACO1, ACLS, ACHE, ASIC2, ACADSB, ABCA4, ABCA3, AMY1A, AMY1C, ART1, AQP5, ARSL, ARSB, ARRB1, PHOX2A, ARHGAP6, ARHGAP1, ARF6, FASLG, AOC2, APOH, APOC1, APOA4, BIRC3, APEH, APCS, APBB1, AKR1C4, CHGB, CHRM2, DSC3, EIF4E, EFNB2, EFNB1, LPAR1, E2F1, DUSP2, RCAN1, DPAGT1, ENO3, DNTT, DNMT3A, SARDH, DLG4, DIO1, DES, DBI, EMD, EPHA3, CHRM3, F8, FGF10, FDXR, FCGR2B, FBLN2, FANCD2, FABP4, F10, F2RL1, EPHA4, EYA3, ETV5, ESRRG, ERCC3, ERBB2, EPOR, EPHX1, DAG1, DAB2, CYP27B1, CCR7, COL9A1, COL8A1, COL4A2, COL1A2, CNP, CNN2, ABCC2, CCR3, CYP24A1, CLCN7, CLCN3, CLC, CISH, CIRBP, CHRNA4, CHRNA3, COL9A2, COL9A3, COMP, KLF6, CYP7A1, CYP4B1, CYP3A7, CYP2A6, CTSK, CTSG, CTRL, CTF1, CTBP1, CSF3, CRYZ, CREBBP, CLDN7, COX4I1, SLC31A1, ODC1, OPRD1, PSD4, CRLF1, SLC22A8, SLC22A6, TRIP10, NREP, KLF4, ASIC3, GPR37L1, MSC, TGFBRAP1, MTA2, XPR1, SLC28A2, HGS, P2RX6, RGN, USP2, COX5A, NTN1, PKD2L1, ARHGEF10, FIG4, RB1CC1, TOX, SLK, PPP6R2, GREB1, PHF14, ABCG1, NCR1, CARTPT, NPEPPS, ADAMTS1, ADAMTS4, HOMER1, GSTO1, MED23, ANGPTL1, NAT8, RABGAP1L, TAM, RGS20, TNFSF11, FCN3, AGPS, SEMA7A, FOXN1, AXIN2, LAP, TP63, PRRC2A, ST8SIA4, SCG2, IL1R2, NPHS2, YY1, XRCC1, PSMG1, AKR1C3, SOCS3, CCN4, F2RL3, KALRN, TRPA1, MGAM, WASF1, PER2, ALKBH1, TMEM11, IRS2, NRP1, HESX1, IL18RAP, TNFRSF10B, TNFSF14, RNGTT, MBTPS1, WASHC5, PUM3, OPRK1, MLXIP, KDM1A, FAIM2, PDCD11, SCAP, SIRT2, SACM1L, CARD8, ZNF652, TBC1D9, PTGDR2, POLI, CHEK2, RASSF1, SLC2A6, WDHD1, CORO1A, NCDN, ZCCHC14, PDIA5, TRS-AGA2-3, ATP6V0A2, RBFOX2, MMD, TRAM1, NPTXR, HEY1, SF3B3, MLYCD, LPIN1, SIRT5, WWC1, ADGRL3, CLEC16A, TASOR, NBEAL2, RRS1, YWHAQ, BTG3, RBM8A, CEBPZ, RAMP3, ABCC4, MSLN, GDF11, ANGPTL7, CALCRL, TSHZ1, BCAP31, BET1, PTPRU, NR1H3, HDAC5, SRA1, SLC12A6, FGFBP1, MVP, STUB1, APBB3, GPR75, DCTN6, FGL2, HPSE, WASF3, GJB6, AHCYL1, KDM5B, TNFSF13B, TXNIP, AKR1A1, HEXIM1, P3H4, ZNRD2, APPBP2, STK25, TFG, TLR6, XPC, NSD2, LAT2, PROS1, PTGDR, TAS2R38, PTBP1, PYY, PSMB8, PSG5, PSEN2, PRNP, PTPRO, PRKCG, PRH2, PRH1, SRGN, PRF1, PPP6C, PPP1CB, PTPRD, PXN, PPIB, RLN1, RRAS, RRAD, RPS6, RPL13, BRD2, RMRP, RLN2, RIEG2, RAC2, RHO, RGS1, REG1A, RBM4, RASA1, RAP1B, RAP1A, PPL, CTSA, VTN, PAM, PDE2A, PCYT1A, PCOS1, PCNA, PC, PAX5, CNTN3, PAK3, PDR, PAK1, P4HB, P2RY2, P2RX4, P2RX1, OTC, OPRM1, PDPK1, PEE1, POU5F1, PLAU, PON2, PODXL, EXOSC10, PLOD1, PLD2, PLCG1, PLCB3, PLAG1, PFKFB3, PLA2G5, PLA2G4A, PLA2G1B, PKM, PIN1, SERPINA1, PHB, S100A1, SORT1, SAA1, TBXA2R, TERT, TEK, TDO2, TCOF1, TCN2, TCF7L2, TBX3, TBX5, TG, TAP1, TAGLN, SYT1, SULT1A3, STAT5A, STAT1, TRIM21, TFRC, THRA, SAA2, UBTF, VIPR2, VDAC3, VDAC2, VDAC1, VAV2, VASP, UGCG, TTN, TIMP2, PHLDA2, TRV-AAC1-4, TRIO, HSP90B2P, TPH1, TPD52, TNFAIP3, SRY, SREBF1, SPRR2A, SDHC, SLC5A1, SLC4A1, SLC3A2, ST3GAL4, SI, SGCA, SEMA3F, SDC1, SPARC, CXCL5, CCL22, CCL13, CCL11, CCL7, SCN8A, SCD, SMTN, SLC6A9, SLC16A1, SLC19A1, SP1, SOS1, SORL1, SORD, SNX1, FSCN1, SNAP25, SMS, SMO, SMN2, SMN1, HLTF, SLC22A5, SLC22A4, SLC22A3, H3P23
-
Bipolar Disorder Not Otherwise Specified
Wikipedia
Bipolar disorder NOS Specialty Psychiatry Bipolar disorder not otherwise specified ( BD-NOS ) is a diagnosis for bipolar disorder (BD) when it does not fall within the other established sub-types. Bipolar disorder NOS is sometimes referred to as subthreshold bipolar disorder. [1] Contents 1 Classification 2 Diagnosis 3 Treatment 4 Epidemiology 5 References 6 External links Classification [ edit ] BD-NOS is a mood disorder and one of three subtypes on the bipolar spectrum, which also includes bipolar I disorder and bipolar II disorder . [1] BD-NOS was a classification in the DSM-IV and has since been changed to Bipolar "Other Specified" and "Unspecified" in the 2013 released DSM-5 (American Psychiatric Association, 2013). ... Bipolar NOS may be diagnosed when it is difficult to tell whether bipolar is the primary disorder due to another general medical condition, such as a substance use disorder. [3] Treatment [ edit ] Individual approaches to treatment are recommended, usually involving a combination of mood stabilizers and atypical antipsychotics . [4] Psychotherapy may be beneficial and should be started early. [4] Epidemiology [ edit ] The prevalence of BD-NOS is 1.4%. [1] References [ edit ] ^ a b c "International impact of bipolar disorder highlights need for recognition and better treatment availability" .
-
Down Syndrome
Wikipedia
They are typically used in combination to increase the detection rate. [20] None can be definitive, thus if screening is positive, either amniocentesis or chorionic villus sampling is required to confirm the diagnosis. [80] Screening in both the first and second trimesters is better than just screening in the first trimester. [80] The different screening techniques in use are able to pick up 90–95% of cases, with a false-positive rate of 2–5%. [81] If Down syndrome occurs in one in 500 pregnancies and the test used has a 5% false-positive rate, this means, of 26 women who test positive on screening, only one will have Down syndrome confirmed. [81] If the screening test has a 2% false-positive rate, this means one of eleven who test positive on screening have a fetus with Down syndrome. [81] First- and second-trimester screening [80] Screen Week of pregnancy when performed Detection rate False positive Description Combined test 10–13.5 wks 82–87% 5% Uses ultrasound to measure nuchal translucency in addition to blood tests for free or total beta-hCG and PAPP-A Quad screen 15–20 wks 81% 5% Measures the maternal serum alpha-fetoprotein, unconjugated estriol, hCG, and inhibin -A Integrated test 15–20 wks 94–96% 5% Is a combination of the quad screen, PAPP-A, and NT Cell-free fetal DNA From 10 wks [86] 96–100% [87] 0.3% [88] A blood sample is taken from the mother by venipuncture and is sent for DNA analysis. ... Findings that indicate increased risk when seen at 14 to 24 weeks of gestation include a small or no nasal bone, large ventricles , nuchal fold thickness , and an abnormal right subclavian artery , among others. [89] The presence or absence of many markers is more accurate. [89] Increased fetal nuchal translucency (NT) indicates an increased risk of Down syndrome picking up 75–80% of cases and being falsely positive in 6%. [90] Ultrasound of fetus with Down syndrome showing a large bladder Enlarged NT and absent nasal bone in a fetus at 11 weeks with Down syndrome Blood tests Several blood markers can be measured to predict the risk of Down syndrome during the first or second trimester. [81] [91] Testing in both trimesters is sometimes recommended and test results are often combined with ultrasound results. [81] In the second trimester, often two or three tests are used in combination with two or three of: α-fetoprotein , unconjugated estriol, total hCG, and free βhCG detecting about 60–70% of cases. [91] Testing of the mother's blood for fetal DNA is being studied and appears promising in the first trimester. [87] [92] The International Society for Prenatal Diagnosis considers it a reasonable screening option for those women whose pregnancies are at a high risk for trisomy 21. [93] Accuracy has been reported at 98.6% in the first trimester of pregnancy. [20] Confirmatory testing by invasive techniques (amniocentesis, CVS) is still required to confirm the screening result. [93] Management Efforts such as early childhood intervention , screening for common problems, medical treatment where indicated, a good family environment, and work-related training can improve the development of children with Down syndrome. ... "Down's syndrome". Lancet (Review). 361 (9365): 1281–89. doi : 10.1016/S0140-6736(03)12987-X . ... PMID 25530442 . ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai Hickey, F; Hickey, E; Summar, KL (2012). ... New York: Demos Medical Pub. p. Chapter 67. ISBN 978-1-934559-86-4 . Archived from the original on 2017-01-23.S100B, SLC19A1, MTHFR, GATA1, RCAN1, SOD1, MIR155, DCR, MIR802, PRDX2, MIR125B2, MIR99A, VIP, PRDX6, CXCL8, CALCA, MIRLET7C, NTF3, GSTM2, DYRK1A, RPLP0, CBS, ETS2, ERG, JAK2, DSCAM, MAPT, CRLF2, MTR, MTRR, RUNX1, TP53, PAPPA, ACTB, APP, CBSL, APOE, AFP, SIM2, RFC1, PSEN1, COL6A1, TTC3, HTC2, CGB3, TNF, S100A1, CGA, POTEF, BACE1, BDNF, SOAT1, KCNJ6, CCL4L1, CCL4, CCL4L2, PLAC4, CGB5, CGB8, COL6A2, IL1B, IL6, CRELD1, DNMT3B, MECP2, SOD2, OLIG2, BACE2, ETV6, PGF, CASP3, RET, HLCS, GET1, EDNRB, PCNT, HMGN1, NEFL, IFNA1, SERPINA1, IFNG, IL1A, DOP1B, SYNJ1, IFNA13, MCIDAS, CAT, AIRE, ABL1, XPNPEP1, SH3BGR, CD6, SQSTM1, CHDH, ITSN1, ENO2, PRNP, SLC25A1, ADAM12, BHMT, IL10, CD19, POLB, CD34, MIR1246, DICER1, ITGB2, EZH2, FABP7, PTCH1, VEGFA, PIGP, GFAP, DNMT3L, CBR1, GART, GAPDH, RAB5A, ALB, HLA-A, PABPC4, TNFRSF10C, CHAF1B, AKT1, SUCLA2, CCL2, SET, FANCB, JAK3, REST, PCP4, BACH1, BCL2, CSF2, MTHFD1, BCHE, LEP, NOS3, CDKN2A, COL18A1, UBB, LAMC2, SNX27, HT, KIT, NFATC1, NGF, USP25, OXA1L, RENBP, NFIC, ABCG1, USP16, PRDX1, NFIB, CCND1, PTGS2, HMOX1, HMGB1, MX1, CALB2, NOP53, NCAM2, NFIA, MMP9, TRIB1, IKZF1, DSCR4, MPL, PIK3CD, KMT2A, APOA1, OPN1SW, BCR, ABCB1, ITGAM, ITGA2B, TARDBP, PKNOX1, INHA, ADIPOQ, RASSF1, PML, PFKL, IL10RB, IL2RA, IL2, IL1RN, IMMT, IGFBP7, STIP1, PRDX3, IFNAR2, RELN, TP73, IFIT3, CFAP410, NAE1, BCAR1, GRIK1, NFIX, GJB2, DHFR, U2AF1, CTNNB1, CTNND1, SYP, DBN1, DCK, DLX4, XRCC1, CD63, DPYSL2, SORL1, CD59, TMX2-CTNND1, MYL7, CSTB, TCN2, CSF3, CSE1L, TYMS, H3P7, TWIST1, ERVH48-1, TTN, P2RY8, COMT, TSHR, TGFB1, TGM2, ESCO1, TNFRSF1A, TPO, SLC12A5, CDA, FLT3, RHD, FAM3B, GATA4, FN1, TREM2, MTOR, PICALM, ARHGEF5, TRRAP, MATR3, USP9X, TXN, VAMP8, PSG10P, POU5F1P4, TNFSF10, TP63, TYRP1, GATD3A, APOBEC3B, MIR590, CCR2, GDF15, TTR, MIR3197, AKR7A2, RCAN2, LOH19CR1, DENR, H3P9, TPTE, HSP90B1, TRH, TRPM2, TSC2, MIR486-1, FXR2, ACTR2, PDXK, DNM1L, POU5F1P3, MAGI2, TMEM72, ZEB2, SCRN1, PHYHIP, ADAMTS1, YWHAG, DSCAM-AS1, GATD3B, EIF2B4, ZMYM2, IL1RL1, EIF2B2, RNF103-CHMP3, AVSD1, LPAR2, UBA3, FAS-AS1, MAP3K14, WT1, XIST, ZFY, HAP1, BCL10, EIF2S2, MTRNR2L12, COX5A, RECQL4, ZNF587B, NTN1, EIF2AK3, MIR3196, NRIP1, ABCG2, UBE3A, USP5, SOD2-OT1, UTRN, UVRAG, USP7, LINC02605, VIPR1, BEST1, MIR1973, MCM3AP, PAX8, VPREB1, TET3, SDS, DCAF7, ACKR3, ENOSF1, KIF21A, SEPTIN11, PARD3, NPDC1, SLC12A9, PLSCR4, STRBP, ACTR3B, AICDA, LRRC47, NUFIP2, MRTFA, RNF213, MIA3, RTL1, DCTN4, RBM11, NT5C3A, PRRX2, TDP2, CHMP3, RIPPLY3, BRWD1, SETD4, PACC1, RIPK4, C21orf91, RBFOX1, TET2, PGPEP1, STX17, LSM2, PRM3, HPSE2, TAF8, AP5B1, UBXN11, SLC13A5, MYL12B, PRRT2, DSCR9, AHSA2P, PRDM16, VENTXP1, LYPD4, STRC, TMPRSS6, ARX, HEPACAM, BAGE2, TMEM241, GLIS2, COL25A1, FGFBP2, ARHGAP24, MED25, LPAL2, TET1, DERL1, TAS2R62P, NANOS3, WNK1, GORASP1, CPEB1, NOD2, SAMSN1, HSPA14, COPS4, EFS, CKAP4, NES, ARPP19, PPARGC1A, PSG8, SUB1, PRSS21, DHFR2, CXCR6, STMN2, MMRN1, MCF2L, ICOSLG, SYNM, ANGPTL2, CHL1, KHDRBS1, APH1A, SLC9A6, TUBB3, RACK1, PEMT, OLFM1, MCRS1, MAD2L2, PLF, MYL12A, CIB1, DPYSL4, MRPL28, GNLY, AHSA1, SPAG5, HEY2, TMEM131, MORC3, REM1, CRCP, APEX2, HPGDS, MIR145, SGSM3, IGHV1-12, NXT1, FAM215A, LGALS13, CNOT7, GMPPA, ZBTB21, IL22, GMNN, GPR162, B3GAT1, CKAP2, PLA2G2D, APPL1, MIR146A, CHD5, MIR17, MTHFD1L, MIR183, MIR191, POC1A, MIR199B, MIR30C1, NUP62, MIR30C2, DAPK2, A1BG, PTH, TNNT2, EDNRA, EEF1A2, EGFR, EGR2, EGR3, EIF2B1, ELANE, ELK1, ENG, ENSA, EPAS1, ERBB2, ERCC2, ERCC3, ERF, ESR1, EEF1A1, E2F1, HP, DSG1, NKX2-5, CYP2B6, CYP17A1, CYP19A1, DBH, DBP, DGCR, DMD, DNAH8, DNASE1, DNMT1, DNMT3A, DPYSL3, DRD4, ATN1, ESR2, FABP3, FABP5, FBN2, GPT, GPX1, GPX3, GSN, GTF2H1, GYPA, GYPB, GYPE, HSD17B10, HAS2, CFH, HINT1, HLA-B, HLA-DQA1, HNMT, GPR42, GPI, GJA1, FLT1, FCGR3A, FCGR3B, FEB1, FGFR1, FGFR3, FLNA, FMR1, GHR, FYN, GABPA, GAD1, GAP43, GDI2, GH1, CST3, CRP, CRMP1, BTG1, ARNT, ARSA, STS, ARSD, ASPA, SERPINC1, AZGP1, BAX, BCL2L1, BCL6, BDKRB1, BGN, BLVRA, BMI1, BRS3, AQP4, APOC2, APOA2, ADRA1A, SERPINA3, ACO1, ACVR1, ADAR, PARP1, PARP4, ADRA2B, APEX1, ADRB2, AGA, AGTR1, AHSG, AMPH, ANK1, BST2, C9, CREB1, CA2, CHRNA7, CHRNB2, CLU, CCR5, CNR1, COL4A3, COL6A3, COL7A1, COX8A, CP, CPE, CPOX, CPS1, CPT1B, CR2, CHRNA4, CHRNA3, CHAT, CDK1, CAMP, CASP1, RUNX1T1, CD247, CD14, CD38, CDK6, CENPB, CDKN1A, CDKN1B, CDKN1C, CDKN2B, CEBPE, CECR, HNRNPA2B1, PRMT2, TLR4, PSMA5, PTPN4, PTPN11, PTPRF, PWP2, RAB4A, RARA, RBM4, RBP4, RCN1, RELA, BRD2, RPE65, RPL17, RPS19, RYR2, A2M, PSG5, HSPA4, PSEN2, PDYN, PGD, PIP4K2A, PLG, PLEK, PLTP, PRRX1, POLD1, PON1, POU5F1, PPP2CA, MAPK10, PRL, TMPRSS15, PSD, SETMAR, SFPQ, SRSF6, SH3BGRL, CDKL5, SYT1, TBX1, TCP1, TRBV20OR9-2, TEAD4, TERC, TFAM, TFCP2, TFF1, TG, THBS1, TIAM1, TIMP3, TLR2, HSPA13, STATH, STAT3, SLC6A4, SHBG, SHMT1, SHMT2, SHOX, SIM1, SLC5A3, SNAP25, STAT1, SNCA, FSCN1, SNRNP70, SOX11, SRY, SSTR4, PDGFB, PDE9A, PDE4C, MDM4, KCNE1, KCNJ15, KISS1, KNG1, KRT8, STMN1, LBP, LBR, LHCGR, LRP2, LTA4H, MAOA, MARK1, MBP, CD46, KCNA3, ITGAL, ISL1, IL7, HSPD1, ICAM1, IDE, IFNAR1, IGH, IL4, IL15, ISG20, IL16, IL17A, IDO1, INPP5D, INSL4, IRF1, MDM2, MME, PDE4B, MOS, NFKB1, NHS, NME1, NME2, NOS2, NPM1, NT5E, NTRK1, NR4A2, OCA2, PAH, PAK1, PAK3, PAX5, PCYT1A, NFE2L2, NFATC2, NEFH, RNR2, MPO, MRC1, MT2A, ATP6, ND3, MTNR1B, MYD88, NDUFV3, MYO9B, NACA, NAIP, NCAM1, NDUFA2, NDUFS3, H3P10
-
Chronic Traumatic Encephalopathy
Wikipedia
Nevertheless, autoimmune changes in blood of players may consist the earliest measurable event predicting CTE. [23] According to 2017 study on brains of deceased gridiron football players, 99% of tested brains of NFL players, 88% of CFL players, 64% of semi-professional players, 91% of college football players, and 21% of high school football players had various stages of CTE. ... Concussion or mild traumatic brain injury (mTBI) is one of the most common neurologic disorders accounting for approximately 90% of all brain injuries sustained (Saulle M, Greenwald BD, 2012). [41] CTE is common among the athletic population because of the high prevalence of concussions; “estimated 1.6–3.8 million sport-related concussion annually in the USA” (Saulle M, Greenwald BD, 2012). [41] This number could be higher because many people do not report their symptoms or may not even know they could have a concussion. ... “In a 2009 review of CTE, McKee et al. found that 51 neuropathologically diagnosed cases of CTE 46 (90%) occurred in athletes. Specifically, athletes participating in American football, boxing, soccer, and hockey comprise the majority of cases” (Saulle M, Greenwald BD, 2012). [41] It is unclear the incidence of CTE as well as at what age it can start. ... PMID 31215252 . ^ Saulle M, Greenwald BD (2012). "Chronic traumatic encephalopathy: a review" (PDF) . ... Molecular and Cellular Neuroscience . 66 : 75–80. doi : 10.1016/j.mcn.2015.03.001 .
-
Systemic Inflammatory Response Syndrome
Wikipedia
Pediatric Critical Care Medicine . 6 (1): 2–8. doi : 10.1097/01.PCC.0000149131.72248.E6 . PMID 15636651 . ^ Goldstein, Brahm; Giroir, Brett; Randolph, Adrienne; International Consensus Conference on Pediatric Sepsis (January 2005). ... Pediatric Critical Care Medicine . 6 (1): 2–8. doi : 10.1097/01.PCC.0000149131.72248.E6 . ISSN 1529-7535 . PMID 15636651 . ^ "Systemic Inflammatory Response Syndrome Treatment & Management" . ... Critical Care Medicine . 35 (9 Suppl): S584–90. doi : 10.1097/01.CCM.0000279189.81529.C4 . ... Current Drug Targets . 10 (9): 872–80. doi : 10.2174/138945009789108774 . ... "An argument for Vitamin E supplementation in the management of systemic inflammatory response syndrome". Shock . 19 (2): 99–103. doi : 10.1097/00024382-200302000-00001 .TNFRSF1B, NOS2, IL6, TNF, CRP, IL10, IL1B, ALB, BTBD8, TLR4, CXCL8, RIPK1, MBL2, HMGB1, TLR2, HGF, HMOX1, LCN2, FCGR1A, IFNG, MASP2, RIPK3, CYBB, CD14, CCL2, THBD, SERPINE1, SCN10A, DCAF1, SELE, SELL, ZFYVE9, SELPLG, SRSF5, SPATA2, PPP6R2, CIR1, SLPI, SPG7, TNFAIP3, HGS, SPP1, ARTN, NRP1, SST, TCF21, SERPINB7, FCN3, TAM, LAT2, VIP, EDIL3, UROD, TNNI3, NR1I3, ADM, TRIM13, STIM2, COLEC11, CORO7, ASRGL1, ZBP1, RPAIN, CHRFAM7A, IL33, SPECC1, NLRP3, SLC18B1, STON1-GTF2A1L, ANKRD42, MIR143, MIR15A, MIR191, MIR23B, CXADRP1, TLX1NB, LOC100505909, NOD2, RTN4, CXCL13, NAT10, SH2B2, CCL27, RSC1A1, GTF2A1L, STON1, CARD8, SIRT2, ARC, PADI4, CLEC5A, LY96, ATRNL1, C5AR2, PDLIM3, CD274, IL19, IRAK4, GDE1, TLR9, RTN1, PIP, RRBP1, CPN1, CXADR, ATN1, EDA, EDN1, ESR1, F2, F3, FKBP1A, FKBP1AP1, FKBP1AP2, FKBP1AP3, FKBP1AP4, FPR1, GABPA, NR3C1, GRP, GSN, CSF2, CISH, CFH, CHI3L1, AFM, AGRP, ALDH2, ALPP, BIRC2, KLK3, ARR3, ATHS, BPI, C5AR1, CALCA, CAPN1, CARS1, CASP1, CASR, CD28, CDK9, HARS1, HP, PTX3, MIF, NFE2L2, NFKB1, NPPB, OTC, PF4, SERPINA1, PIK3CA, PIK3CB, PIK3CD, PIK3CG, ADRB2, PLG, PPBP, PPIA, PRKAR1A, PTEN, PTH, COX1, MEFV, AGFG1, MC1R, HRG, HSPA1A, HSPA1B, HSPA4, HSP90AA1, DNAJC4, ICAM1, IL1A, IL1RN, IL4, CXCR1, ITGB1, ITGB2, KCNJ5, LAMC2, LEP, LTA, MIR4772
-
Cross-Reactivity
Wikipedia
Journal of Analytical Toxicology . 38 (7): 387–96. doi : 10.1093/jat/bku075 . PMID 24986836 . ^ Porrozzi R, Teva A, Amaral VF, Santos da Costa MV, Grimaldi G (September 2004). ... PMID 24950742 . ^ Kasprowicz V, Ward SM, Turner A, Grammatikos A, Nolan BE, Lewis-Ximenez L, Sharp C, Woodruff J, Fleming VM, Sims S, Walker BD, Sewell AK, Lauer GM, Klenerman P (March 2008). ... The Journal of Allergy and Clinical Immunology . 102 (6 Pt 1): 1005–12. doi : 10.1016/S0091-6749(98)70339-2 . PMID 9847442 . ^ Altmann F (2007). ... International Archives of Allergy and Immunology . 142 (2): 99–115. doi : 10.1159/000096114 . PMID 17033195 .
-
Hughes–stovin Syndrome
Wikipedia
The syndrome is also identified as being "characterized by pulmonary/bronchial artery aneurysms and thrombophlebitis, without diagnostic features of Behçet's disease (BD)." [4] Management [ edit ] There is currently no satisfactory treatment for this condition. [4] This is partly due to its rarity; the Orphanet Journal of Rare Diseases claims that fewer than forty cases of the disease have been described in English medical literature. [5] Immunosuppressive therapy is the most common treatment, involving a mix of glucocorticoids and cyclophosphamide . ... "Hughes-Stovin Syndrome: a case report and review of the literature" . Cases J . 2 : 98. doi : 10.1186/1757-1626-2-98 . PMC 2649053 .
-
Perineal Tear
Wikipedia
American Journal of Obstetrics and Gynecology . 200 (5): 573.e1–573.e7. doi : 10.1016/j.ajog.2008.11.022 . ... "Antenatal perineal massage to prevent birth trauma". American Family Physician . 89 (5): 335–6. PMID 24695503 . ^ Wang, H; Jayasekara, R; Warland, J (12 March 2015). ... Third degree obstetric anal sphincter tears: risk factors and outcome of primary repair" . BMJ . 308 (6933): 887–91. doi : 10.1136/bmj.308.6933.887 . ... Lancet . 354 (9183): 983–986. doi : 10.1016/S0140-6736(98)11205-9 . PMID 10501360 . S2CID 37825406 . ^ Moore JE, Witt WP, Elixhauser A (April 2014). ... Obstetrics and Gynecology . 107 (6): 1261–8. doi : 10.1097/01.aog.0000218693.24144.bd . PMID 16738150 . S2CID 23901136 . ^ Lammers, K; Prokop, M; Vierhout, ME; Kluivers, KB; Fütterer, JJ (August 2013).
-
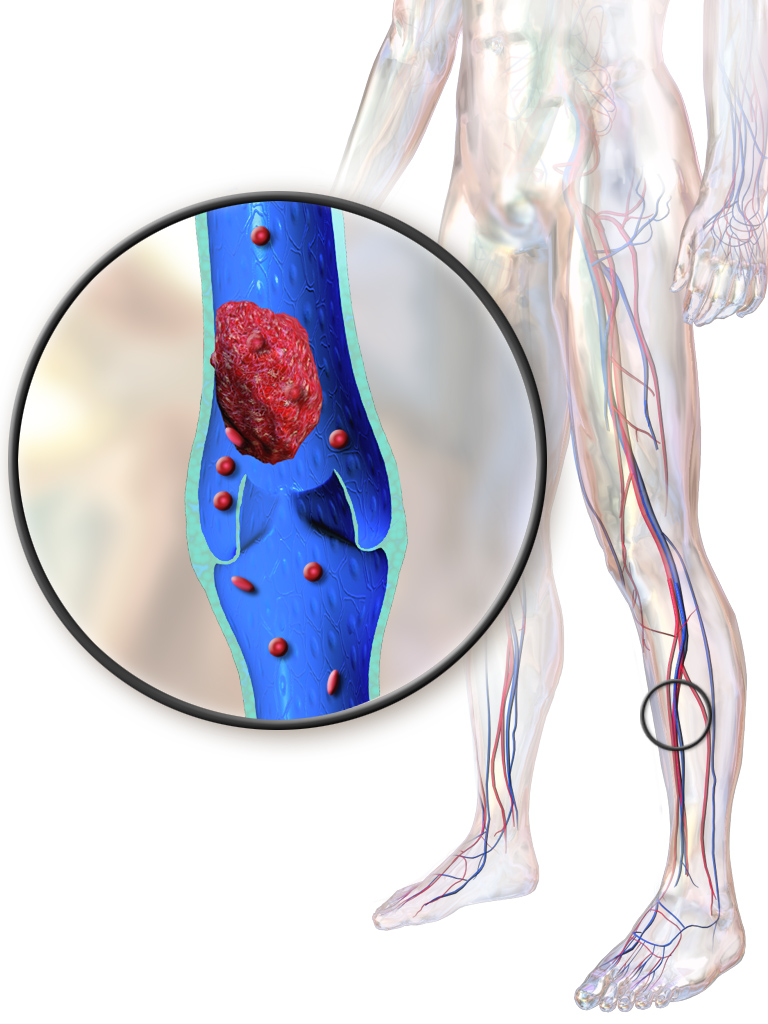
Hughes-Stovin Syndrome
Orphanet
Hughes-Stovin syndrome (HSS) is a life-threatening disorder, believed to be a cardiovascular clinical variant manifestation of Behçet's disease (BD; see this term). It is characterized by the association of multiple pulmonary artery aneurysms (PAAs) and peripheral venous thrombosis. ... Etiology The etiology of HSS is unknown; however, it is assumed that HSS is a form of vasculitis following a similar mechanism of pathogenesis to that thought to be involved in BD. Diagnostic methods Diagnosis of HSS is made on the basis of the clinical picture (association of venous thrombosis and PAAs in a young patient), patient history and imaging studies (chest radiographs, conventional angiography or helical computed tomography) for detection and evaluation of the PAAs. ... Differential diagnosis The pulmonary manifestations of HSS and BD have been reported to be identical, but the two syndromes can be distinguished on the basis of the absence of mucocutaneous findings in HSS.
-
Trigonocephaly
Wikipedia
Optima Grafische Communicatie. ISBN 978-90-8559-601-1 . ^ Wilkie, AO (1997). ... The Journal of Pediatrics . 93 (2): 188–91. doi : 10.1016/S0022-3476(78)80493-4 . ... Plastic and Reconstructive Surgery . 97 (2): 276–81. doi : 10.1097/00006534-199602000-00002 . ... Plastic and Reconstructive Surgery . 80 (2): 195–212. doi : 10.1097/00006534-198708000-00006 . ... American Journal of Medical Genetics Part A . 146A (8): 984–91. doi : 10.1002/ajmg.a.32208 . PMID 18344207 .
-
Iatrogenic Calcinosis Cutis
Wikipedia
New England Journal of Medicine . 356 (6): e5. doi : 10.1056/NEJMicm055763 . PMID 17287472 .
- Subungual Exostosis Wikipedia
-
Complement Component 4a Deficiency
Omim
Associated characteristics indicated a defect in synthesis of C4 and a genetic basis thereof was indicated by the fact that C4 phenotyping in 20 patients and in 26 parents showed that 90% and 81%, respectively, had null allotypes at either the C4A or C4B (120820) locus compared with 59% in controls. ... They confirmed the earlier findings of high frequencies of C4-null phenotypes and of HLA-B8,DR3 antigens. In addition, they found that among the patients most of both the C4A- and C4B-null phenotypes resulted from gene deletions.