-
Epileptic Encephalopathy, Early Infantile, 27
Omim
These patients were identified through several research settings focusing on neurodevelopmental disorders. Variants in 86 of 91 patients were classified as putatively pathogenic; all 86 patients presented with developmental delay, intellectual disability (ID), and/or autism spectrum disorder (ASD). ... The mutations were found by targeted massive parallel resequencing of 50 known EIEE genes plus candidate genes in 357 patients with epilepsy, including 91 patients with epileptic encephalopathy. The patients bearing mutations thus accounted for 2.2% (2 of 91) of that phenotypic group. In vitro functional expression studies showed that both mutations occurred in the ion channel-forming reentrant loop and resulted in increased calcium permeability and a gain of function.CDKL5, ARX, STXBP1, TSC2, SPTAN1, SCN2A, GRIN2B, ST3GAL3, CNPY3, PLCB1, PHACTR1, SIK1, PIGA, POMC, GUF1, NTRK2, MAGI2, TSC1, CRH, WDR45, HSD17B4, UPB1, AIMP2, TBCD, ABAT, CFL1, ND1, ALG13, SLC35A2, CASK, NEUROD2, TRNW, TRNV, TRNL1, TRNK, PNKP, ND6, ND5, PIGQ, ND4, ND3, ND2, ZNHIT3, ATP6, HIBCH, PIGP, KIF1A, RFT1, CLCN4, KCNA1, TRIM8, SLC19A3, NGLY1, IFNG, TUBA8, SLC25A22, GNAO1, KCNQ2, CRHR1, GABRB3, HTC2, MECP2, FOXG1, NR3C1, MC2R, KCNJ6, NR2F1, PAFAH1B1, SCN1A, PIGW, HNRNPU, TBC1D24, IARS2, BRWD3, HDAC4, KCNT2, KPNA7, DEPDC5, WDR62, AKT3, CPP, PRRT2, UBA5, PDCD6IP, ARFGEF2, TIMM50, FARS2, ATG10, TLK2, SORCS3, TBL1XR1, KCNT1, ARC, PARS2, RARS2, MBD5, PNPO, AARS1, PTCH1, KCNAB2, DLX5, GNB1, GABRB2, GABRA1, MTOR, FLNA, FGF12, FBN1, DPYS, DNMT3A, DNM1, DLG3, YWHAG, DCX, CTNNB1, CSNK1E, CHD2, CACNA2D1, C5AR1, C5, BTD, APC, ACTB, GRIN1, GRIN2A, IGF1, IL2RG, TWIST1, TCOF1, SPTBN2, SOS1, SLC6A4, SLC1A4, SKI, SCN3A, ATXN2, RASGRF1, PPP1CB, PMM2, ABCB1, PDHA1, NGF, NDP, MEF2C, MC4R, LAMA2, KCNJ11, ITGA2B, LINC02210-CRHR1
-
Levator Ani Syndrome
Wikipedia
ISBN 978-88-470-1542-5 . ^ a b c Bharucha AE, Trabuco E (September 2008). "Functional and chronic anorectal and pelvic pain disorders" . ... PMID 15526112 . ^ Rao, SS; Bharucha, AE; Chiarioni, G; Felt-Bersma, R; Knowles, C; Malcolm, A; Wald, A (25 March 2016). ... "Anorectal and Pelvic Pain" . Mayo Clinic Proceedings (review). 91 (10): 1471–1486. doi : 10.1016/j.mayocp.2016.08.011 .
-
Waldenström's Macroglobulinemia
Wikipedia
A skeletal survey can help distinguish between WM and multiple myeloma . [48] Anemia is typically found in 80% of patients with WM. A low white blood cell count , and low platelet count in the blood may be observed. ... Retrieved 24 September 2020 . ^ a b c d Cheson BD (2006). "Chronic Lymphoid Leukemias and Plasma Cell Disorders". ... "Living with Waldenstrom's Macroglobulinemia." ^ Kyle RA (1998). "Chapter 94: Multiple Myeloma and the Dysproteinemias". ... Leuk Res . 15 (6): 407–8. doi : 10.1016/0145-2126(91)90049-Y . PMID 1907339 . ^ Baehring, J.; Hochberg, E.; Raje, N.; Ulrickson, M.; Hochberg, F. (2008). ... "Characterization of subpopulation lacking both B-cell and plasma cell markers in Waldenstrom macroglobulinemia cell line" . Lab. Invest . 94 (1): 79–88. doi : 10.1038/labinvest.2013.129 .MYD88, PAX5, IGH, LPL, CXCR4, ANPEP, MYC, ZAP70, MALT1, CDR3, MYOM2, SDC1, CXCL12, AIMP2, TCL1A, GRAP2, TCL1B, AHSA1, ACSBG1, RNF19A, IBTK, POLDIP2, IGHV4-34, BCL11A, KRT20, EXOC2, RAPH1, MAPK1, NOS1, NOTCH2, NOS2, BCL6, BCL9, BLM, BRAF, BSG, BTK, SERPING1, CCND3, CD6, MS4A1, TNFRSF8, CD40LG, CD79A, CRK, MAPK14, CTNNB1, IL4, IRF4, CD200, NCAM1, CCND1, SLC35B2
-
Oral Mite Anaphylaxis
Wikipedia
The World Allergy Organization Journal . 2 (5): 91–6. doi : 10.1186/1939-4551-2-5-91 .
-
Nephrotoxicity
Wikipedia
Normal creatinine level is between 80 - 120 μmol/L. In interventional radiology, a patient's creatinine clearance levels are all checked prior to a procedure. ... Nature Clinical Practice Nephrology . 2 (2): 80–91. doi : 10.1038/ncpneph0076 . PMID 16932399 .
-
Progressive Nonfluent Aphasia
Wikipedia
Neurol . 49 (4): 425–32. doi : 10.1002/ana.91 . PMID 11310619 . ^ Gorno-Tempini ML, Hillis AE, Weintraub S, et al. ... Curr Neurol Neurosci Rep . 10 (6): 484–90. doi : 10.1007/s11910-010-0140-4 . ... PMID 14561797 . Reilly J, Rodriguez AD, Lamy M, Neils-Strunjas J (2010).
-
Abortion In Australia
Wikipedia
Before then abortion law was for many years governed by case law under sections 82–84 of the Crimes Act 1900 of New South Wales . ... The offence is called "killing unborn child" and can be committed only around the time of childbirth [80] in Queensland , [81] Western Australia , [82] and the Northern Territory . [83] It is called "causing death of child before birth" in Tasmania . [84] In South Australia, it comes under the heading of " abortion ". [85] The definition is somewhat broader in the Australian Capital Territory , [80] [86] and comparably broad to English law in Tasmania [84] and South Australia . [85] [80] The offence was abolished in Victoria by the Abortion Law Reform Act 2008 (Victoria) . [87] New South Wales does not have a child destruction enactment, [80] but the Crimes Amendment (Grievous Bodily Harm) Act 2005 (NSW) amended the Crimes Act 1900 (NSW) so that s 4(1)(a) now defines "grievous bodily harm" as including "the destruction (other than in the course of a medical procedure) of the foetus of a pregnant woman, whether or not the woman suffers any other harm". [88] This was further amended by the Abortion Law Reform Act 2019 to "the destruction (other than in the course of a medical procedure or a termination of a pregnancy in accordance with the Abortion Law Reform Act 2019 ) of the foetus of a pregnant woman, whether or not the woman suffers any other harm." ... Less than 2% took place at or after 20 weeks. [89] Public opinion [ edit ] Main article: Societal attitudes towards abortion Since at least the 1980s, opinion polls have shown a majority of Australians support abortion rights, [90] and that support for abortion is increasing. [91] In 1987, a Saulwick poll found only about 7 per cent of Australians would not approve of abortions under any circumstances. [92] In 2003, a poll by The Australian Survey of Social Attitudes (AuSSA) found that 81% of Australians believe a woman should have the right to choose an abortion, and 9% believe they should not; 7% were neutral and 2% could not decide. [91] In 2005, a Nielsen Corporation poll found that 56% of Australians thought the abortion laws in place, which generally allow abortion for the sake of life, health, or economic factors, were "about right", 16% want changes in law to make abortion "more accessible" and 17% want changes to make it "less accessible". [93] In 2006, a poll by Roy Morgan Research found that 65% of the Australians approved of surgical abortion and 22% disapproved, and that 62% believed RU-486 should be available to women while 31% believed it should not. [94] In 2006, interviews found that 80% of Australians disapproved the use of sex-selective abortion . [95] In 2007, a poll by AuSSA found that 4% of Australians are opposed to abortion in all circumstances, 33% believe abortion should be allowed in certain circumstances and 57% believed it should be readily available whenever a woman wants one; 7% were undecided or did not respond. [96] In 2009, a study of polls conducted during Australia's 2007 federal elections found that a clear majority of both Labor Party and Liberal Party voters support abortion rights. [96] The study also showed that 77 per cent of winning candidates in the 2007 election favoured an unrestricted approach to abortion. [97] In 2010, a study published in The Medical Journal of Australia found that 61% of Australians said abortion should be lawful without question in the first trimester of pregnancy, while 26% said it should be lawful depending on the reason. [98] In the second trimester and third trimesters, support for outright lawful abortion was 12% and 6% respectively, while 57% and 42% respectively said it depended on the circumstances. [99] See also [ edit ] Abortion in New Zealand Abortion law Child destruction References [ edit ] ^ a b "Abortion Law in Australia" . ... Archived from the original on 27 December 2019. ^ R v Davidson (Menhennitt ruling) [1969] VicRp 85 , [1969] VR 667 (3 June 1969), Supreme Court (Vic, Australia). ^ a b c R v Wald (1971) 3 NSW DCR 25. ... Criminal Code Act Compilation Act 1913 (PDF) (v14-b0-05 ed.). 27 June 2009. p. 142 . Retrieved 31 March 2009 . ^ "Criminal Code Act – Notes" .
-
Chronic Testicular Pain
Wikipedia
"The aetiology, pathophysiology and management of chronic orchialgia" . BJU . 91 (5): 435–437. doi : 10.1046/j.1464-410x.2003.04094.x . ... "Clinical, radiographic, and pathologic description of IgG4-related perivasal fibrosis: a previously undescribed etiology of chronic orchialgia" . Urology . 84 (4): 748–750. doi : 10.1016/j.urology.2014.06.035 .
-
Neonatal Hypoglycemia
Wikipedia
One way of screening includes a heel stick to test the blood glucose level at the bedside. [1] Diagnosing hypoglycemia in neonates requires two consecutive blood glucose readings of less than 40 mg/dl to properly diagnose hypoglycemia. [1] Bedside glucose monitoring is only effective if the equipment is accurate, rapid, and reliable. [1] This form of testing is often faster and more cost effective. [1] Laboratory serum glucose testing is the most accurate way of testing blood glucose levels. [1] These specimens are either taken from the heel, arterial, or venous punctures and must be store immediately on ice in order to prevent glycolysis, further altering the results. [1] USA guidelines recommended that the hypoglycemic neonate should have a glucose test every 2–4 hours for the first 24 hours of life. [1] Guidelines in the UK, however, recommend pre-feed screening of at-risk infants at 2–4 hours of age (to avoid false positives when blood glucose is, ordinarily, at its lowest at 2–3 hours of age) and at the subsequent feed until a blood glucose level of >2.0 mmol/l (36 mg/dl) on at least two consecutive occasions and is feeding well. [ citation needed ] Management [ edit ] Some infants are treated with 40% dextrose (a form of sugar) gel applied directly to the infant's mouth. [10] There are two main ways that neonatal hypoglycemia is treated. [1] The first way includes intravenous infusion of glucose. [1] For less severe, borderline, asymptomatic hypoglycemic neonates early introduction of breast milk can be effective for raising glucose levels to a healthy level. [1] Any infant at risk of hypoglycemia should have their blood sugar taken again one hour after birth. [1] Oral glucose is another option to restore normal glucose levels if the newborn is having difficulty latching to the breast or breastfeeding is not an option, however, breast milk is proven to be a better source as it includes glucose and carbohydrates. [1] It is recommended by The American Academy of Pediatrics that infants feed within the first hour of life with the glucose reading being 30 minutes after this feeding for an accurate result. [1] If the initial feeding does not raise the newborn's blood glucose above 40 mg/dl then the newborn must receive an IV infusion of 10% dextrose in water as a mini bolus as 2 ml/kg over 1 minute. [1] Following the mini bolus a continuous infusion of 10% dextrose in water at 80-100 ml/kg/day in order to maintain a healthy serum glucose level between 40–50 mg/dl. [1] Maintaining newborn thermoregulation is a large part in preventing further hypoglycemia as well. [1] Nursing care management [ edit ] The biggest nursing concern for a neonate experiencing hypoglycemia is the physical assessment to potentially find the cause. [1] It is also essential to prevent environmental factors such as cold stress that may predispose the newborn for further decreasing blood sugar. [1] Within the physical assessment, comorbidities of hypoglycemia should also be assessed such as intolerance of feeding, or respiratory distress. [1] Another important nursing intervention is assisting the mother in successful breastfeeding as this can prevent and treat hypoglycemia. [1] If an IV infusion of 10% dextrose in water is initiated then the nurse must monitor for: •Circulatory overload [1] •Hyperglycemia [1] •Glycosuria [1] •Intracellular dehydration [1] See also [ edit ] Congenital hyperinsulinism Hyperinsulinemic hypoglycemia References [ edit ] ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al am an ao ap aq ar as at au av aw ax ay az ba bb bc bd be bf bg bh bi bj bk bl bm bn bo Wong's nursing care of infants and children .HNF4A, HNF1A, TBX19, GCK, AKT2, ZMPSTE24, PROP1, ABCC8, UCP2, KDM6A, KLF11, PIGQ, TRIP11, KCNQ1OT1, NEUROD1, APPL1, WAC, POMP, PRKAG2, ASXL2, GATAD2B, NSD1, CLPB, DNAJC19, H19, PAX4, H19-ICR, GYS2, GALC, FBP1, GPC3, KCNQ1, HADH, DBH, CYP11A1, HSD3B2, IGF2, CEL, INS, PDX1, CDKN1C, KCNJ11, BLK, ATP5F1D, MTOR, ALDH7A1, GH1, MLYCD, HIF1A, FOXA2, IGFBP3, HLTF, POMC, PCK1, NHS
-
Internal Hernia
Wikipedia
"Über Darmverschlinung nach der Gastroenterostomie". Arch Klin Chir . 62 : 94–114. ^ Cho, M; Pinto, D; Carrodeguas, L; Lascano, C; Soto, F; Whipple, O; Simpfendorfer, C; Gonzalvo, JP; Zundel, N (2006). ... Surgery for Obesity and Related Diseases . 2 (2): 87–91. doi : 10.1016/j.soard.2005.11.004 .
-
Eosinophilic Folliculitis
Wikipedia
Eosinophilic folliculitis Other names Eosinophilic pustular folliculitis, Sterile eosinophilic pustulosis" [1] Specialty Dermatology Eosinophilic folliculitis is an itchy rash with an unknown cause that is most common among individuals with HIV , though it can occur in HIV-negative individuals where it is known by the eponym Ofuji disease . [2] EF consists of itchy red bumps ( papules ) centered on hair follicles and typically found on the upper body, sparing the abdomen and legs. ... Contents 1 Pathophysiology 2 Diagnosis 3 Treatment 4 Epidemiology 5 See also 6 References 7 External links Pathophysiology [ edit ] The cause of EF is unknown. A variety of microorganisms have been implicated, including the mite Demodex , [3] the yeast Pityrosporum , [4] and bacteria . [5] An autoimmune process has what is this also been investigated. [6] Diagnosis [ edit ] Eosinophilic folliculitis may be suspected clinically when an individual with HIV exhibits the classic symptoms. ... Treatment [ edit ] Treatment of eosinophilic folliculitis in people with HIV typically begins with the initiation of Highly Active Anti-Retroviral Therapy in order to help reconstitute the immune system. Direct treatment of the EF itself focuses on decreasing the inflammation and itching. ... Other therapies include PUVA , topical tacrolimus , and isotretinoin . [7] Epidemiology [ edit ] Eosinophilic folliculitis associated with HIV infection typically affects individuals with advanced HIV and low T helper cell counts. [8] It affects both men and women as well as children with HIV and is found throughout the world. EF may also affect individuals with hematologic disease such as leukemia and lymphoma . [9] It may also affect otherwise normal infants in a self-limited form. [10] HIV-negative individuals can also develop EF — this is more common in Japan. [11] See also [ edit ] Eosinophilic pustular folliculitis of infancy List of cutaneous conditions References [ edit ] ^ James, William; Berger, Timothy; Elston, Dirk (2005). ... Archives of Dermatology . 131 (9): 1089–91. doi : 10.1001/archderm.131.9.1089 .
-
Vacterl Association
Wikipedia
Please help improve this article by adding citations to reliable sources . ... Contents 1 Signs and symptoms 1.1 Spine 1.2 Anal defects 1.3 Cardiac defects 1.4 Trachea and esophagus 1.5 Kidneys 1.6 Limbs 1.7 Extension 1.8 Growth 2 Pathology 3 Diagnosis 3.1 Differential diagnosis 4 Management 5 Epidemiology 6 History 7 See also 8 References 9 Further reading 10 External links Signs and symptoms [ edit ] The following features are observed with VACTERL association: [2] [3] V - Vertebral anomalies A - Anorectal malformations C - Cardiovascular anomalies T - Tracheoesophageal fistula E - Esophageal atresia R - Renal (Kidney) and/or radial anomalies L - Limb defects Although it was not conclusive whether VACTERL should be defined by at least two or three component defects, [4] it is typically defined by the presence of at least three of the above congenital malformations. [5] Spine [ edit ] Vertebral anomalies, or defects of the spinal column, usually consist of small ( hypoplastic ) vertebrae or hemivertebra where only one half of the bone is formed. About 80 percent of patients with VACTERL association will have vertebral anomalies. [6] In early life these rarely cause any difficulties, although the presence of these defects on a chest x-ray may alert the physician to other defects associated with VACTERL. ... Babies with VACTERL association, however, do tend to have normal development and normal intelligence. [ citation needed ] Pathology [ edit ] Patients with abnormal cardiac and kidney function may be more at risk for hemolytic uremic syndrome Diagnosis [ edit ] Differential diagnosis [ edit ] Baller-Gerold syndrome [5] CHARGE syndrome [5] Currarino syndrome [5] DiGeorge Syndrome [5] Fanconi anemia [5] Feingold syndrome [5] Fryns syndrome [5] MURCS association [5] Oculo-auriculo-vertebral syndrome [5] Opitz G/BBB syndrome [5] Pallister–Hall syndrome [5] Townes–Brocks syndrome [5] VACTERL with hydrocephalus [5] [8] Management [ edit ] This section is empty. You can help by adding to it . ( December 2017 ) Epidemiology [ edit ] The incidence of VACTERL association is estimated to be approximately 1 in 10,000 to 1 in 40,000 live-born infants. [5] It is seen more frequently in infants born to diabetic mothers. ... PMID 8826430 . ^ a b c d e f g h i j k l m n o Solomon, BD (Aug 16, 2011). "VACTERL/VATER Association" .
-
Anti-Iglon5 Disease
Wikipedia
Treatment [ edit ] Anti-IgLON5 disease is mainly treated with immunosuppressants (80%), mostly cycles of IV corticosteroids (58%) in combination with IV immunoglobulins (IVIg−36%) and/or TPE (27%). ... "A Proteomic Screen of Neuronal Cell-Surface Molecules Reveals IgLONs as Structurally Conserved Interaction Modules at the Synapse" . Structure . 27 (6): 893–906.e9. doi : 10.1016/j.str.2019.03.004 . ... "IgLON5: A case with predominant cerebellar tau deposits and leptomeningeal inflammation". Neurology . 91 (4): 180–182. doi : 10.1212/WNL.0000000000005859 .
-
Down Syndrome
Wikipedia
They are typically used in combination to increase the detection rate. [20] None can be definitive, thus if screening is positive, either amniocentesis or chorionic villus sampling is required to confirm the diagnosis. [80] Screening in both the first and second trimesters is better than just screening in the first trimester. [80] The different screening techniques in use are able to pick up 90–95% of cases, with a false-positive rate of 2–5%. [81] If Down syndrome occurs in one in 500 pregnancies and the test used has a 5% false-positive rate, this means, of 26 women who test positive on screening, only one will have Down syndrome confirmed. [81] If the screening test has a 2% false-positive rate, this means one of eleven who test positive on screening have a fetus with Down syndrome. [81] First- and second-trimester screening [80] Screen Week of pregnancy when performed Detection rate False positive Description Combined test 10–13.5 wks 82–87% 5% Uses ultrasound to measure nuchal translucency in addition to blood tests for free or total beta-hCG and PAPP-A Quad screen 15–20 wks 81% 5% Measures the maternal serum alpha-fetoprotein, unconjugated estriol, hCG, and inhibin -A Integrated test 15–20 wks 94–96% 5% Is a combination of the quad screen, PAPP-A, and NT Cell-free fetal DNA From 10 wks [86] 96–100% [87] 0.3% [88] A blood sample is taken from the mother by venipuncture and is sent for DNA analysis. ... Findings that indicate increased risk when seen at 14 to 24 weeks of gestation include a small or no nasal bone, large ventricles , nuchal fold thickness , and an abnormal right subclavian artery , among others. [89] The presence or absence of many markers is more accurate. [89] Increased fetal nuchal translucency (NT) indicates an increased risk of Down syndrome picking up 75–80% of cases and being falsely positive in 6%. [90] Ultrasound of fetus with Down syndrome showing a large bladder Enlarged NT and absent nasal bone in a fetus at 11 weeks with Down syndrome Blood tests Several blood markers can be measured to predict the risk of Down syndrome during the first or second trimester. [81] [91] Testing in both trimesters is sometimes recommended and test results are often combined with ultrasound results. [81] In the second trimester, often two or three tests are used in combination with two or three of: α-fetoprotein , unconjugated estriol, total hCG, and free βhCG detecting about 60–70% of cases. [91] Testing of the mother's blood for fetal DNA is being studied and appears promising in the first trimester. [87] [92] The International Society for Prenatal Diagnosis considers it a reasonable screening option for those women whose pregnancies are at a high risk for trisomy 21. [93] Accuracy has been reported at 98.6% in the first trimester of pregnancy. [20] Confirmatory testing by invasive techniques (amniocentesis, CVS) is still required to confirm the screening result. [93] Management Efforts such as early childhood intervention , screening for common problems, medical treatment where indicated, a good family environment, and work-related training can improve the development of children with Down syndrome. ... Retrieved 20 March 2012 . ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad Malt, EA; Dahl, RC; Haugsand, TM; Ulvestad, IH; Emilsen, NM; Hansen, B; Cardenas, YE; Skøld, RO; Thorsen, AT; Davidsen, EM (Feb 5, 2013). ... Tidsskrift for den Norske Laegeforening . 133 (3): 290–94. doi : 10.4045/tidsskr.12.0390 . PMID 23381164 . ^ a b c d e f g h i j k l m n o Kliegma, Robert M. (2011). ... PMID 25530442 . ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai Hickey, F; Hickey, E; Summar, KL (2012).S100B, SLC19A1, MTHFR, GATA1, RCAN1, SOD1, MIR155, DCR, MIR802, PRDX2, MIR125B2, MIR99A, VIP, PRDX6, CXCL8, CALCA, MIRLET7C, NTF3, GSTM2, DYRK1A, RPLP0, CBS, ETS2, ERG, JAK2, DSCAM, MAPT, CRLF2, MTR, MTRR, RUNX1, TP53, PAPPA, ACTB, APP, CBSL, APOE, AFP, SIM2, RFC1, PSEN1, COL6A1, TTC3, HTC2, CGB3, TNF, S100A1, CGA, POTEF, BACE1, BDNF, SOAT1, KCNJ6, CCL4L1, CCL4, CCL4L2, PLAC4, CGB5, CGB8, COL6A2, IL1B, IL6, CRELD1, DNMT3B, MECP2, SOD2, OLIG2, BACE2, ETV6, PGF, CASP3, RET, HLCS, GET1, EDNRB, PCNT, HMGN1, NEFL, IFNA1, SERPINA1, IFNG, IL1A, DOP1B, SYNJ1, IFNA13, MCIDAS, CAT, AIRE, ABL1, XPNPEP1, SH3BGR, CD6, SQSTM1, CHDH, ITSN1, ENO2, PRNP, SLC25A1, ADAM12, BHMT, IL10, CD19, POLB, CD34, MIR1246, DICER1, ITGB2, EZH2, FABP7, PTCH1, VEGFA, PIGP, GFAP, DNMT3L, CBR1, GART, GAPDH, RAB5A, ALB, HLA-A, PABPC4, TNFRSF10C, CHAF1B, AKT1, SUCLA2, CCL2, SET, FANCB, JAK3, REST, PCP4, BACH1, BCL2, CSF2, MTHFD1, BCHE, LEP, NOS3, CDKN2A, COL18A1, UBB, LAMC2, SNX27, HT, KIT, NFATC1, NGF, USP25, OXA1L, RENBP, NFIC, ABCG1, USP16, PRDX1, NFIB, CCND1, PTGS2, HMOX1, HMGB1, MX1, CALB2, NOP53, NCAM2, NFIA, MMP9, TRIB1, IKZF1, DSCR4, MPL, PIK3CD, KMT2A, APOA1, OPN1SW, BCR, ABCB1, ITGAM, ITGA2B, TARDBP, PKNOX1, INHA, ADIPOQ, RASSF1, PML, PFKL, IL10RB, IL2RA, IL2, IL1RN, IMMT, IGFBP7, STIP1, PRDX3, IFNAR2, RELN, TP73, IFIT3, CFAP410, NAE1, BCAR1, GRIK1, NFIX, GJB2, DHFR, U2AF1, CTNNB1, CTNND1, SYP, DBN1, DCK, DLX4, XRCC1, CD63, DPYSL2, SORL1, CD59, TMX2-CTNND1, MYL7, CSTB, TCN2, CSF3, CSE1L, TYMS, H3P7, TWIST1, ERVH48-1, TTN, P2RY8, COMT, TSHR, TGFB1, TGM2, ESCO1, TNFRSF1A, TPO, SLC12A5, CDA, FLT3, RHD, FAM3B, GATA4, FN1, TREM2, MTOR, PICALM, ARHGEF5, TRRAP, MATR3, USP9X, TXN, VAMP8, PSG10P, POU5F1P4, TNFSF10, TP63, TYRP1, GATD3A, APOBEC3B, MIR590, CCR2, GDF15, TTR, MIR3197, AKR7A2, RCAN2, LOH19CR1, DENR, H3P9, TPTE, HSP90B1, TRH, TRPM2, TSC2, MIR486-1, FXR2, ACTR2, PDXK, DNM1L, POU5F1P3, MAGI2, TMEM72, ZEB2, SCRN1, PHYHIP, ADAMTS1, YWHAG, DSCAM-AS1, GATD3B, EIF2B4, ZMYM2, IL1RL1, EIF2B2, RNF103-CHMP3, AVSD1, LPAR2, UBA3, FAS-AS1, MAP3K14, WT1, XIST, ZFY, HAP1, BCL10, EIF2S2, MTRNR2L12, COX5A, RECQL4, ZNF587B, NTN1, EIF2AK3, MIR3196, NRIP1, ABCG2, UBE3A, USP5, SOD2-OT1, UTRN, UVRAG, USP7, LINC02605, VIPR1, BEST1, MIR1973, MCM3AP, PAX8, VPREB1, TET3, SDS, DCAF7, ACKR3, ENOSF1, KIF21A, SEPTIN11, PARD3, NPDC1, SLC12A9, PLSCR4, STRBP, ACTR3B, AICDA, LRRC47, NUFIP2, MRTFA, RNF213, MIA3, RTL1, DCTN4, RBM11, NT5C3A, PRRX2, TDP2, CHMP3, RIPPLY3, BRWD1, SETD4, PACC1, RIPK4, C21orf91, RBFOX1, TET2, PGPEP1, STX17, LSM2, PRM3, HPSE2, TAF8, AP5B1, UBXN11, SLC13A5, MYL12B, PRRT2, DSCR9, AHSA2P, PRDM16, VENTXP1, LYPD4, STRC, TMPRSS6, ARX, HEPACAM, BAGE2, TMEM241, GLIS2, COL25A1, FGFBP2, ARHGAP24, MED25, LPAL2, TET1, DERL1, TAS2R62P, NANOS3, WNK1, GORASP1, CPEB1, NOD2, SAMSN1, HSPA14, COPS4, EFS, CKAP4, NES, ARPP19, PPARGC1A, PSG8, SUB1, PRSS21, DHFR2, CXCR6, STMN2, MMRN1, MCF2L, ICOSLG, SYNM, ANGPTL2, CHL1, KHDRBS1, APH1A, SLC9A6, TUBB3, RACK1, PEMT, OLFM1, MCRS1, MAD2L2, PLF, MYL12A, CIB1, DPYSL4, MRPL28, GNLY, AHSA1, SPAG5, HEY2, TMEM131, MORC3, REM1, CRCP, APEX2, HPGDS, MIR145, SGSM3, IGHV1-12, NXT1, FAM215A, LGALS13, CNOT7, GMPPA, ZBTB21, IL22, GMNN, GPR162, B3GAT1, CKAP2, PLA2G2D, APPL1, MIR146A, CHD5, MIR17, MTHFD1L, MIR183, MIR191, POC1A, MIR199B, MIR30C1, NUP62, MIR30C2, DAPK2, A1BG, PTH, TNNT2, EDNRA, EEF1A2, EGFR, EGR2, EGR3, EIF2B1, ELANE, ELK1, ENG, ENSA, EPAS1, ERBB2, ERCC2, ERCC3, ERF, ESR1, EEF1A1, E2F1, HP, DSG1, NKX2-5, CYP2B6, CYP17A1, CYP19A1, DBH, DBP, DGCR, DMD, DNAH8, DNASE1, DNMT1, DNMT3A, DPYSL3, DRD4, ATN1, ESR2, FABP3, FABP5, FBN2, GPT, GPX1, GPX3, GSN, GTF2H1, GYPA, GYPB, GYPE, HSD17B10, HAS2, CFH, HINT1, HLA-B, HLA-DQA1, HNMT, GPR42, GPI, GJA1, FLT1, FCGR3A, FCGR3B, FEB1, FGFR1, FGFR3, FLNA, FMR1, GHR, FYN, GABPA, GAD1, GAP43, GDI2, GH1, CST3, CRP, CRMP1, BTG1, ARNT, ARSA, STS, ARSD, ASPA, SERPINC1, AZGP1, BAX, BCL2L1, BCL6, BDKRB1, BGN, BLVRA, BMI1, BRS3, AQP4, APOC2, APOA2, ADRA1A, SERPINA3, ACO1, ACVR1, ADAR, PARP1, PARP4, ADRA2B, APEX1, ADRB2, AGA, AGTR1, AHSG, AMPH, ANK1, BST2, C9, CREB1, CA2, CHRNA7, CHRNB2, CLU, CCR5, CNR1, COL4A3, COL6A3, COL7A1, COX8A, CP, CPE, CPOX, CPS1, CPT1B, CR2, CHRNA4, CHRNA3, CHAT, CDK1, CAMP, CASP1, RUNX1T1, CD247, CD14, CD38, CDK6, CENPB, CDKN1A, CDKN1B, CDKN1C, CDKN2B, CEBPE, CECR, HNRNPA2B1, PRMT2, TLR4, PSMA5, PTPN4, PTPN11, PTPRF, PWP2, RAB4A, RARA, RBM4, RBP4, RCN1, RELA, BRD2, RPE65, RPL17, RPS19, RYR2, A2M, PSG5, HSPA4, PSEN2, PDYN, PGD, PIP4K2A, PLG, PLEK, PLTP, PRRX1, POLD1, PON1, POU5F1, PPP2CA, MAPK10, PRL, TMPRSS15, PSD, SETMAR, SFPQ, SRSF6, SH3BGRL, CDKL5, SYT1, TBX1, TCP1, TRBV20OR9-2, TEAD4, TERC, TFAM, TFCP2, TFF1, TG, THBS1, TIAM1, TIMP3, TLR2, HSPA13, STATH, STAT3, SLC6A4, SHBG, SHMT1, SHMT2, SHOX, SIM1, SLC5A3, SNAP25, STAT1, SNCA, FSCN1, SNRNP70, SOX11, SRY, SSTR4, PDGFB, PDE9A, PDE4C, MDM4, KCNE1, KCNJ15, KISS1, KNG1, KRT8, STMN1, LBP, LBR, LHCGR, LRP2, LTA4H, MAOA, MARK1, MBP, CD46, KCNA3, ITGAL, ISL1, IL7, HSPD1, ICAM1, IDE, IFNAR1, IGH, IL4, IL15, ISG20, IL16, IL17A, IDO1, INPP5D, INSL4, IRF1, MDM2, MME, PDE4B, MOS, NFKB1, NHS, NME1, NME2, NOS2, NPM1, NT5E, NTRK1, NR4A2, OCA2, PAH, PAK1, PAK3, PAX5, PCYT1A, NFE2L2, NFATC2, NEFH, RNR2, MPO, MRC1, MT2A, ATP6, ND3, MTNR1B, MYD88, NDUFV3, MYO9B, NACA, NAIP, NCAM1, NDUFA2, NDUFS3, H3P10
-
Scott Syndrome
Wikipedia
Blood 1996; 87:1409-1415 ^ Elliott JI, Mumford AD, Albrecht C, Collins PW, Giddings JC, Higgins CF et al. Characterization of lymphocyte responses to Ca2+ in Scott syndrome. Thromb Haemost 2004; 91:412-415 ^ Sims PJ, Wiedmer T. Unraveling the mysteries of phospholipid scrambling. ... Production and characterization of transformed B-lymphocytes expressing the membrane defect of Scott Syndrome. J Clin Invest 1994; 94:2237-2244 Stout JG, Basse F, Luhm RA, Weiss HJ, Wiedmer T, Sims PJ. Scott syndrome erythrocytes contain a membrane protein capable of mediating Ca2+-dependent transbilayer migration of membrane phospholipids. J Clin Invest 1997; 99:2232-2238 Albrecht C, McVey JH, Elliott JI, Sardini A, Kasza I, Mumford AD, et al. ... A hereditary bleeding disorder of dogs caused by a lack of platelet procoagulant activity. Blood 2002; 99:2434-2441 External links [ edit ] Classification D ICD - 10 : D69.8 OMIM : 262890 MeSH : C563120 C563120, C563120 DiseasesDB : 32153 External resources Orphanet : 806
-
Murcs Association
Wikipedia
Please help improve this article by adding links that are relevant to the context within the existing text. ( April 2014 ) ( Learn how and when to remove this template message ) MURCS association Other names Müllerian duct aplasia-renal dysplasia-cervical somite anomalies syndrome This condition can be inherited in an autosomal dominant manner(though not always) [1] Specialty Medical genetics MURCS association (a variant of Mayer-Rokitansky-Küster-Hauser syndrome ) is a very rare developmental disorder [2] that primarily affects the reproductive and urinary systems involving MU llerian agenesis , R enal agenesis , C ervicothoracic S omite abnormalities. [3] It affects only females. Contents 1 Genetics 2 Diagnosis 3 Treatment 4 Notes 5 References 6 External links Genetics [ edit ] Genetic heterogeneity is observed in MURCS association. [4] Diagnosis [ edit ] This section is empty. You can help by adding to it . ( August 2017 ) Treatment [ edit ] This section is empty. You can help by adding to it . ( August 2017 ) Notes [ edit ] ^ RESERVED, INSERM US14 -- ALL RIGHTS. of diseases=Mayer-Rokitansky-Kuster-Hauser-syndrome-type-2&title=Mayer-Rokitansky-Kuster-Hauser-syndrome-type-2&search=Disease_Search_Simple "Orphanet: Mayer Rokitansky Kuster Hauser syndrome type 2" Check |url= value ( help ) . www.orpha.net . ... "MURCS association with additional congenital anomalies". Human Pathology . 17 (1): 88–91. doi : 10.1016/s0046-8177(86)80160-5 .
-
Cowden Syndrome 6
Omim
Molecular Genetics Orloff et al. (2013) found that, of 91 individuals with Cowden syndrome without germline PTEN (601728), SDHB (185470), or SDHD (602690) mutation or KLLN (612105) promoter hypermethylation, 2 individuals (2.2%) carried 1 of 2 germline missense mutations in the AKT1 gene.PTEN, SDHB, PIK3CA, KLLN, AKT1, SDHD, SEC23B, SDHC, FGFR2, EGFR, TNF, NOD2, TEP1, IL10, BMPR1A, TP53, IL23R, OSR1, BRCA1, MTOR, ATG16L1, SDS, CEACAM6, SARDH, CRP, HLA-DQA1, TGM2, IFNG, SMAD4, KLK3, IL17A, BRCA2, ACAD8, SORD, IL6, PLAG1, PIK3CG, NPEPPS, TGFBI, TESC, PIK3CD, PIK3CB, IL1B, PSAT1, NR3C1, GTF2H1, GTF2H2, GTF2H3, GTF2H4, HIF1A, IBD5, TGFB1, MAPK1, HSP90B1, ADA, TLR2, TLR4, STK11, BDNF, SST, MIR21, SLC12A3, GTF2H5, IL15, TPMT, TSC1, LTA, LCN2, TNFSF15, VWF, IL15RA, PROS1, ERCC2, ERCC3, HEPH, DGCR2, DNAJC6, GDF15, GRAP2, CLOCK, TP63, ARHGEF2, HMGA2, TIMP1, TNFRSF1A, TNNI3, TSC2, UBC, VEGFA, VIM, AIMP2, GPR68, SLC16A3, BFSP2, RASAL1, RABEPK, NR1I2, SQSTM1, SGCE, USP8, SLC16A4, ABCB6, ACAT1, LANCL1, SEMA4D, LYPD1, TAGAP, SLCO6A1, SIRPA, SPRED1, TMPRSS6, USF3, IL27, TCERG1L, ARMH1, GSTK1, ASPG, MIR19A, MIR206, MIR29B1, MIR29B2, DEFA1A3, POU5F1P3, POU5F1P4, DEFB4B, MIR1290, LINC01672, OCLN, RN7SL263P, LOC110806263, IL17F, LMLN, KAT8, IL23A, AHSA1, TXNIP, CCR9, EBNA1BP2, KIF3A, FSTL1, TBC1D9, RNF19A, PTPN22, HAVCR1, IL19, MPC1, TMPRSS13, IL17D, SLC38A2, IL26, MYDGF, SUCNR1, SLURP1, HAMP, GORASP1, P2RY12, WNK1, CYREN, POLDIP2, SCD, TG, PTK2B, ACE, DEFB4A, DMBT1, DSC2, DSG2, TOR1A, EGF, ERCC5, ERCC6, EYA1, EYA2, FAP, CYBB, FGF2, FKBP5, FN1, GABPA, GAPDH, GCH1, GH1, GHSR, GLS, FFAR2, GPX1, CYP11A1, CCN2, TFF3, CASP3, ALB, ALDH1A1, APC, AR, ARSD, ATF3, BAX, BCL2, BGLAP, DDR1, CAMP, CAT, CSH2, CAV1, SERPINH1, CCNH, CD86, CD40, CDH2, CDK7, ERCC8, CRK, MAPK14, CSH1, HFE, HLA-DPB1, HLA-DQA2, CCL2, PMS2, POMC, POU5F1, PPIA, MAPK8, PROX1, PSMD7, RAP1A, RELA, RET, ACHE, CCL11, HLA-DQB1, CCL20, SDC1, SLC2A1, SLC6A4, SLC11A1, STAT3, STAT5A, STAT5B, SYT1, ADAM17, TRBV20OR9-2, ABCB1, PGF, TNFRSF11B, NFE2L2, HLA-DRB4, HSPA9, IFNA1, IFNA13, IL4, IL9, JAK2, KCNN4, RPSA, LBR, EPCAM, SMAD2, SMAD7, MAOA, MAT1A, MBL2, MAP3K5, MMP9, MSMB, MST1, MUC4, MUTYH, MYCN, H3P28
-
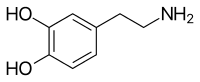
Dopamine Dysregulation Syndrome
Wikipedia
Parkinsonism Relat. Disord . 14 (4): 273–80. doi : 10.1016/j.parkreldis.2007.09.007 . PMID 17988927 . ^ a b c d e f g h i j Lawrence AD, Evans AH, Lees AJ (October 2003). ... The Journal of Neuropsychiatry and Clinical Neurosciences . 26 (3): E3. doi : 10.1176/appi.neuropsych.13060126 . ... "Treatment of cognitive, psychiatric, and affective disorders associated with Parkinson's disease" . Neurotherapeutics . 11 (1): 78–91. doi : 10.1007/s13311-013-0238-x . ... "Impulsive and compulsive behaviors in Parkinson's disease" . Annu Rev Clin Psychol . 10 : 553–80. doi : 10.1146/annurev-clinpsy-032813-153705 .
-
Plaid Syndrome
Wikipedia
JAMA Dermatology ^ Zhou Q, Lee GS, Brady J, Datta S, Katan M, Sheikh A, Martins MS, Bunney TD, Santich BH, Moir S, Kuhns DB, Long Priel DA, Ombrello A, Stone D, Ombrello MJ, Khan J, Milner JD, Kastner DL, Aksentijevich I (2012) A hypermorphic missense mutation in PLCG2, encoding phospholipase Cγ2, causes a dominantly inherited autoinflammatory disease with immunodeficiency. Am J Hum Genet 91(4):713-720. doi: 10.1016/j.ajhg.2012.08.006 ^ Yu P, Constien R, Dear N, Katan M, Hanke P, Bunney TD, Kunder S, Quintanilla-Martinez L, Huffstadt U, Schröder A, Jones NP, Peters T, Fuchs H, de Angelis MH, Nehls M, Grosse J, Wabnitz P, Meyer TP, Yasuda K, Schiemann M, Schneider-Fresenius C, Jagla W, Russ A, Popp A, Josephs M, Marquardt A, Laufs J, Schmittwolf C, Wagner H, Pfeffer K, Mudde GC (2005) Autoimmunity and inflammation due to a gain-of-function mutation in phospholipase C gamma 2 that specifically increases external Ca 2+ entry. ... Front Immunol 9:2863 ^ Ombrello MJ, Remmers EF, Sun G, et al (2012) Cold urticaria, immunodeficiency, and autoimmunity related to PLCG2 deletions.
-
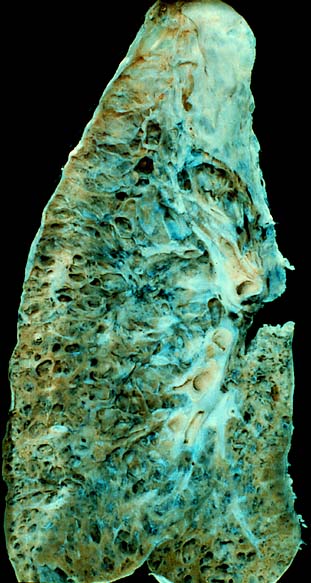
Interstitial Lung Disease
Wikipedia
"European protocols for the diagnosis and initial treatment of interstitial lung disease in children" . review. Thorax . 70 (11): 1078–84. doi : 10.1136/thoraxjnl-2015-207349 . ... The British Journal of Radiology . 90 (1071): 20160723. doi : 10.1259/bjr.20160723 . ... American Journal of Respiratory and Critical Care Medicine . 199 (3): e5–e23. doi : 10.1164/rccm.201812-2276ST . ... PMID 30707039 . ^ Dowman L, Hill CJ, Holland AE (October 2014). "Pulmonary rehabilitation for interstitial lung disease".SFTPC, COPA, FARSB, SOD2, IL18, IL1A, CCL2, MUC5B, SFTPB, HLA-DRB1, ITGA3, STING1, MUC2, NKX2-1, MARS1, TRIM21, ZKSCAN1, WNT3, AP3D1, NSF, SMPD1, SFTPD, AKAP13, MUC1, MAPT, IVD, FAM13A, CTNNA3, ATP11A, LINC02210, LINC02210-CRHR1, TSBP1-AS1, MAPT-AS1, ADAMTS7P4, KRTAP5-4, KANSL1, COPD, RNF168, SPPL2C, LRRC34, DPP9, ERF, DISP2, ACTRT3, STN1, DEPTOR, CSMD1, IFIH1, INPP5E, BTNL2, MYNN, TOLLIP, GBA, ABCA3, LINC02689, SCARB2, DSP, EGFR, CRP, TNF, ALK, PLF, RGS6, CHI3L1, RASA1, SLC27A5, DCPS, CEL, SLURP1, RIEG2, PRTN3, PARP9, RBM45, CCL18, TGFB1, SMUG1, TRIM33, PAH, MORC3, IFNG, MMP7, PADI4, PROS2P, IL17A, SFTPA1, CXCL8, IL2, CD274, IL1B, ADAM17, TERT, CSF2, TNFSF13B, ACR, SFTPA2, CYCS, KRT20, FLI1, PADI3, WNT5A, BLK, CABIN1, ABCA4, TP53, CXCL13, CAV1, FLNA, HARS1, FEV, GPT, TFF1, CRYGD, IL33, MS4A1, CX3CL1, HARS2, ROS1, CXCL10, CDC42EP1, PLEKHM3, HSPA4, ALB, IFNA1, IFNA13, IGF1, BTG3, IL4, IL6, IL10, IRF5, CNTRL, COX8A, AP3B1, LOXL2, TRBV28, TRBV20-1, CXCR6, PSS, MPC1, CD226, ACTB, SCLY, CD40LG, PADI2, SEC14L2, LILRB1, FOXP3, CD53, ERVK-12, IL17RA, PRPF31, ADH5, ERVK-11, CNMD, ERVK-2, BRD4, TRBV3-1, HPGDS, WDHD1, ERVK-22, PSIP1, BICD2, MIR7-1, TLR9, AQP4, HPS4, BCL2, BPIFB1, PIK3IP1, FOXP4, PGA3, C11orf40, IL34, ATF3, AKT1, CD22, SPAAR, FAS, AIRE, MIR7-3, GLIS3, ALOX5AP, ALOX5, MIR7-2, MIR200C, TSLP, BDNF, NT5C1A, SLA2, ADM, BANK1, RNPC3, CD19, CD14, MIR214, NCF1, CXCL16, ACE2, NOD2, CANX, CALR, PINK1, FSD1, KLF5, SLC2A10, MAP1LC3B, PGA4, FSD1L, POSTN, ETS1, YAP1, DNASE1L3, IL1RN, IL15, EGF, ITGAE, ITGAM, SLC26A3, KIT, KRT7, KRT14, LGALS1, LYZ, CYP2B6, NTF3, MATN1, MMP1, MMP9, MPO, MRC1, MSN, MST1, CX3CR1, CTNNB1, MUC5AC, NARS1, IDUA, ICAM2, ICAM1, HSPG2, FCER2, FCGR2B, FCN2, FGF2, FOXF1, FOLR2, FOSL2, ELN, GH1, GLRX, GPR15, GSTM1, HCRT, HDAC2, HGF, HLA-B, HLA-DPB1, EIF2B1, HLA-DRB5, HPS1, HSPA1A, HSPA1B, HSP90AA1, CCN2, PDGFA, TRAIP, EIF2B4, VCAM1, VEGFA, EZR, VTN, VWF, CCR3, FBN1, CXCR4, HMGA2, SEMA7A, FCN3, EIF2B2, PGA5, EIF2S2, LRRFIP1, CD163, PPIG, COX5A, NTN1, ITM2B, CXCL14, CCS, CENPB, PLXNC1, TTF1, TNFRSF1B, TNFRSF1A, COL4A5, PLG, SERPINF2, PSMB2, PTPRC, BRD2, S100A9, SAFB, CSF2RA, CCL3, CCL5, CCL17, CCL21, SELPLG, CPOX, SNRNP70, SPP1, SSTR2, STAT3, STAT4, TAL1, TRB, TGFA, TGFBR1, CHIT1